High resolution cryo-EM images from Africa pave the way to bespoke nitrilases for industry
A team from Cape Town have recently published the first high-resolution cryo-electron microscopy (EM) paper to originate from Africa. As described in Nature Communications Biology, the team solved the structure of a nitrilases enzyme to a close-to-atomic resolution and used the structural insights to design a mutant enzyme that could be fine-tuned for applications in biotechnology. This work was made possible through an access program funded by the Synchrotron Techniques for African Research and Technology project, a collaborative grant which seeks to build partnerships between world leading scientists in Africa and the UK working together on research using synchrotron science.
Nitrilases are a fascinating class of plant enzymes that are significant players in the synthesis of a broad range of important chemicals. These enzymes typically have specificity for a small range of substrates, yet they have huge biotechnological potential. A team of scientists from the University of Cape Town set out to realize this potential by investigating the structure of the enzymes using cryo-EM at the electron Bio-imaging Centre (eBIC).
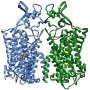
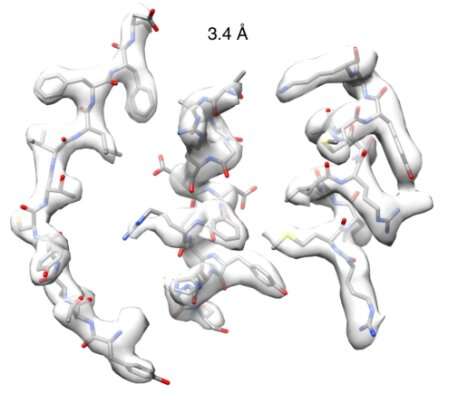
After successfully managing to obtain high-quality structural information, the team obtained a close-to-atomic resolution image (3.4 Å) of a nitrilase derived from the cabbage family. Using this structure, the team semi-rationally designed a new mutant nitrilase that acted on substrates not catalyzed by any other naturally occurring nitrilases.
The team hope that they can progress to making 'designer' nitrilases for any substrate needed by industry—whether pharmaceuticals, fine chemicals or even food. More work at the eBIC is planned to continue this fruitful study.
Helical twists
Nitrilases are a class of enzymes that are used to produce carboxylic acids, ammonia and amides for the large-scale synthesis of medicines and industrially important chemicals. A team of scientists from the University of Cape Town was fascinated by these enzymes and wanted to study how they evolved and correlate their structure and function.
The group initially compared two different nitrilases from Arabidopsis thaliana (a member of the cabbage family) and saw that one had a broad range of substrates, but the other one was much more specific to just a small number of substrates. They carried out a series of mutations to the binding pocket of the enzymes and found out that switching a single amino acid changed the substrate preference of the nitrilases. With this tantalizing knowledge, the group decided obtain detailed structural information to reveal how nitrilases pick their substrates.
Dr. Jeremy Woodward, Lecturer in Medical Biochemistry at the University of Cape Town and Principal Investigator of the study explained his group's motivations,
"To try and figure out what was going on we imaged all the enzymes using low-resolution cryo-EM at the University of Cape Town and found that they formed filaments, and that the tightness of their helical twist was correlated with substrate size. In fact, the amino acid we discovered was located at an interface between two helical subunits. We've observed this correlation with a large number of nitrilase enzymes, but couldn't explain what was happening at the molecular level until our recent visit to eBIC at Diamond Light Source."
Close-to-atomic structure
The team had previously used negative staining electron microscopy (EM) which is limited to a resolution of 20 Å, and is not high enough to see atomic details in the structure. Additionally, prior work from other labs had shown that nitrilases cannot be crystallized, so cryo-EM was their only option.
"This was the only method that could be used to answer the questions that they had," explained Dr. Adriana Klyszejko, post-doctoral research associate at the eBIC who assisted with the study. "At Diamond, what we have created is an integrated effort to our user community where they can push their research further."
At the eBIC, the team obtained a structure at a resolution of 3.4 Å. Dr. Woodward, elaborated:
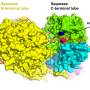
"The high quality of the data we obtained allowed us to visualize the structure of an intact nitrilase helical filament at close-to-atomic resolution for the first time. We observed a loop, held in position by the amino acid we discovered, that limits the maximum size of bound substrates and shifts with helical twist."
The START grant
The work was made possible with a Synchrotron Techniques for African Research and Technology (START) grant; an initiative set up in March 2019 to build partnerships between scientists in Africa and the UK. START is funded by a £3.7M grant by the UKRI's Science and Technology Facilities Council (STFC) from the Global Challenges Research Fund (GCRF). The STFC awarded the funding from the GCRF, a 5-year £1.5Bn fund that is a key component in the delivery of the UK Aid Strategy, ensuring that UK research takes a leading role in addressing the problems faced by developing countries through research and innovation.
The motivation for START comes from the societal challenges faced by African communities; for example, 600 million people (70%) in sub-Saharan Africa live without electricity, and a reliable electricity supply is one of the most powerful tools for lifting people out of poverty and ending their dependency on aid. START researchers will investigate energy materials, including solar cell structures, catalysts and batteries. Development of healthcare in Africa is hampered by a lack of fundamental understanding of the cause in diseases such as malaria or HIV. Structural biology gives unprecedented insight into the mechanisms behind such diseases.
As well as providing access to the world-class facilities at the eBIC to the scientists from South Africa, the grant also supported post-doctoral scientist, Dr. Andani Mulelu who was the lead investigator of the study. "This work would not have been possible without this, because the equipment, support and infrastructure necessary to successfully conduct this experiment is not available in Africa," explained Dr. Woodward.
Dr. Gwyndaf Evans, Principal beamline scientist on Diamond's VMXm beamline, and Life Sciences Principal Investigator for the START project comments:
"The START initiative aims, among other things, to help build expertise and capacity for structural biology in Africa. Initially we are placing emphasis on facilitating access to Diamond's structural biology facilities though focused training courses in South Africa and locally here at Diamond. The ultimate aim is to demonstrate to African funding bodies the power and benefits of structural biology in addressing African problems in human health and agriculture to make structural biology a sustainable activity on the continent. Seeing experts like Jeremy Woodward achieve a tenured position in South Africa bodes well for the future and goes some way to building this sustainability."
Designer nitrilases
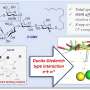
Using the insights gained from the high-resolution cryo-EM structure, the team screened over 5,000 mutants to design a new enzyme with an altered helical twist that acts on a new set of substrates not catalyzed by any other plant nitrilases. This was carried out by identifying 'hotspot' amino acids for directed evolution and selecting them by coupling the survival of bacteria to the successful conversion of a library of substrates.
Building on this work, the team hope to fine-tune nitrilases to realize their full biotechnological potential. "We would like to get to the point where we can produce 'designer' nitrilases for any substrate by making appropriate changes to the helical twist as well as the binding pocket. To achieve this, we would like to visualize a collection of key nitrilases with a range of different helical states (and substrate specificities) by high resolution cryo-EM," concluded Prof Woodward.
More information: Andani E. Mulelu et al. Cryo-EM and directed evolution reveal how Arabidopsis nitrilase specificity is influenced by its quaternary structure, Communications Biology (2019). DOI: 10.1038/s42003-019-0505-4
Journal information: Communications Biology
Provided by Diamond Light Source