March 26, 2019 feature
Cellular microRNA detection with miRacles

MicroRNAs (miRNAs) are short noncoding regulatory RNAs that can repress gene expression post-transcriptionally and are therefore increasingly used as biomarkers of disease. Detecting miRNAs can be arduous and expensive as they require amplification, labelling and radioactive probes. In a recent report published on Science Advances, Arun Richard Chandrasekaran and co-workers at the RNA Institute and Department of Biological Sciences, at the University of Albany, State University of New York, reported on a single-step, non-enzyme microRNA detection assay using conformationally unresponsive DNA nanoswitches.
The scientists named the assay 'miRacles,' to abbreviate 'micro-RNA activated conditional looping of engineered switches'. The assay demonstrated subattomole and single-nucleotide specificity using an agarose gel electrophoresis readout. In the experiments, they detected cellular microRNAs from RNA extractions obtained from differentiating muscles at the nanogram-scale. The scientists presented a cost-effective experimental setup to detect miRNAs in an hours' time frame to provide a compelling alternative to the existing methods of quantitative polymerase chain reaction (qPCR) and Northern blotting to quantify the regulatory genetic materials.
miRNAs can regulate many biological processes during normal physiological development and disease by affecting cell proliferation, differentiation and apoptosis in vivo. The expression of miRNAs can be quantified in tissues, cells and body fluids as stable biomarkers for cellular events and disease diagnosis, highlighting the importance of their detection. Nevertheless, miRNA detection is challenging due to low abundance, small size and similarities in sequence. The biomolecules comprise approximately 0.01 percent of the total RNA content and individual mRNA copies per cell range can widely vary. Additionally, miRNAs in a family can differ by a single nucleotide, while specific miRNAs can be regulated during disease and regular cellular function. As a result, the strategies of detection for miRNA require high specificity and the ability to correctly identify a few molecules from a sample abundant with predominant RNA molecules.
Traditional methods for miRNA detection include Northern blotting, quantitative reverse-transcription polymerase chain reaction (qRT-PCR), next-generation sequencing and microarray based hybridization to separate the miRNA signal from noise. Of the methods specified, Northern blotting can directly identify native miRNAs, while others rely on additional labelling methods or step-wise amplification, adding to the cost, complexity and performance of detection. For example, innovative DNA nanostructures can be used for miRNA detection, where various research groups have previously combined nanostructures with nanoparticles, hybridization chain reactions and transition-metal dichalcogenide nanosheets to enable the process.
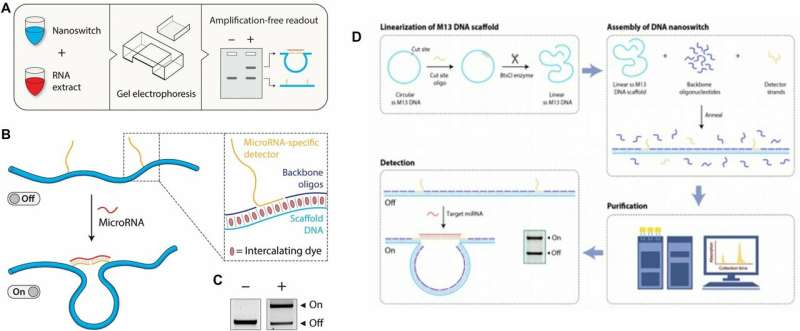
In the present work, Chandrasekaran et al. developed a relatively simple DNA-based device to solve a complex biomedical challenge. In the miRacles assay, the scientists used a 'smart reagent' composed of rationally designed DNA nanoswitches for simple and cost-effective native miRNA detection without using specialized equipment in the lab. DNA nanoswitches were originally designed as tools for single-molecule biophysics experiments and later recognized for their capacity to detect and quantify biomolecular interactions using gel electrophoresis. Previous collaborative research efforts by the same research team focused on molecular detection to quantify protein levels and detect synthetic DNA sequences as a proof-of-concept.
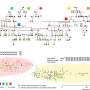
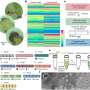
The present work expanded on preliminary studies and concepts to produce user-ready multiplexed miRNA detection and quantification. The scientists analyzed nanograms of cellular RNA extracts in a short time-frame using an experimental setup built using common laboratory supplies. They designed the DNA nanoswitch as a linear duplex that formed a loop in the presence of the target miRNA molecule. To construct the nanoswitch, Chandrasekaran et al. used the DNA origami approach by hybridizing short oligonucleotides complementary to a single-stranded DNA scaffold.
They designed two distant "detector" strands with overhangs complementary to different segments of the target miRNA. When the miRNA recognized and bound to the construct, the switch reconfigured from the linear "off" state to the looped "on" state. They quantified the two states using standard agarose gel electrophoresis to detect the signal arising from the looped nanoswitch. The signal was only amplified by a single miRNA of interest, the results compared favorably with the fluorescence resonance energy transfer (FRET) technique.
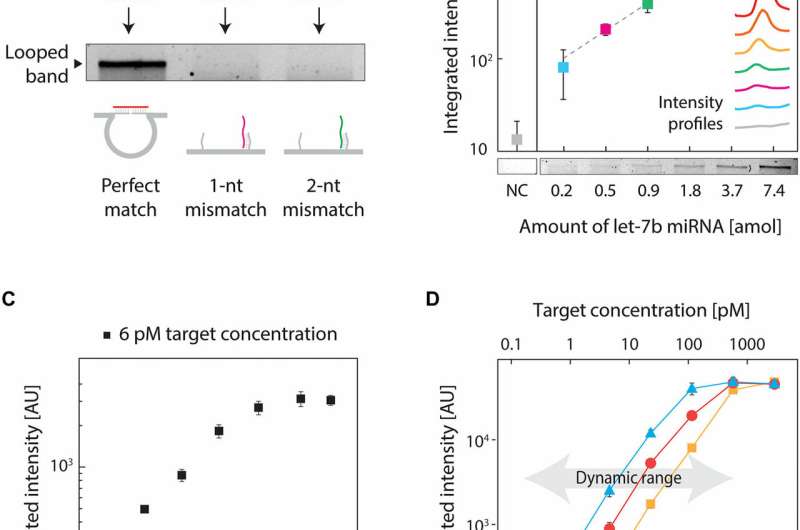
To validate the concept, the team of researchers chose a let-7b target miRNA due to its highly conserved family of more than a dozen-related miRNAs that varied by one or more nucleotides. These miRNAs were suited due to their critical role in biological functions and dysregulation in multiple human diseases. To eliminate cross-talk and noise between the nanoswitch and the target—which led to reduced signal intensity compared to a perfect match, the scientists rationally redesigned the nanoswitches. To achieve perfect specificity, they destabilized the interaction on the side containing the mismatch. Results of the study illustrated high specificity of the assay thus developed, providing an answer to a key challenge in miRNA detection that culminated from a high signal/noise ratio.
The low abundance of miRNAs also required high-sensitivity of detection, which the scientists achieved by optimizing the protocol. They then performed similar experiments for two other variants of miRNAs (miR-15 a and miR-206), resulting in subattomole to single-attomole levels of detection. For instance, for a low concentration of a target sample (6 pM), the signal increased for 4 hours, with little change beyond that timeframe.
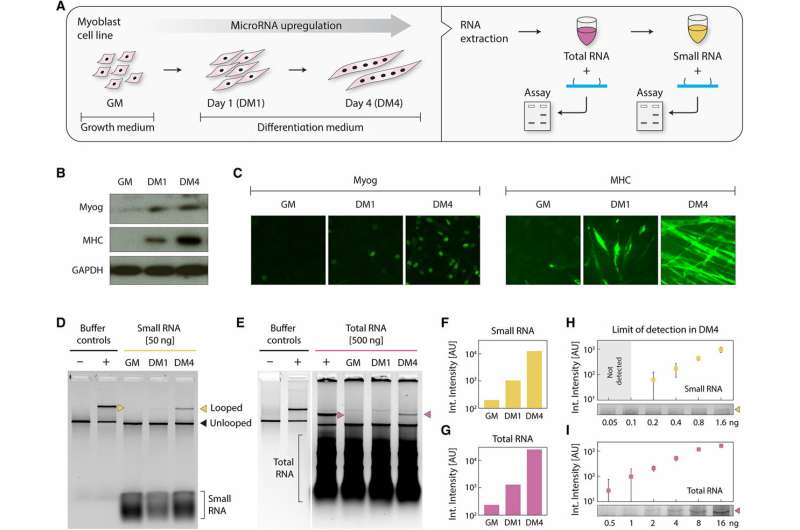
miRNAs are also typically studied in cell cultures, tissue samples and bodily fluids and readily processed to collect total RNAs in the microgram scale, along with a subfraction of small RNA. For this, Chandrasekaran et al. used myoblast cells as a surrogate model system of muscle differentiation, with miR-206 as the primary target. During cell culture, they identified miRNAs in differentiating skeletal muscles, where the molecule miR-206 was significantly upregulated during late differentiation. When the scientists harvested the undifferentiated and differentiated muscle cells to extract small RNAs, they confirmed cell differentiation using Western blots and conducted immunocytochemistry for early and late differentiation markers myogenin (Myog) and myosin heavy chain (MHC) respectively. Since individual miRNAs make up parts-per-million level by mass of total RNAs, detecting miRNAs is as challenging as finding a needle in a haystack. To validate the assay for biological detection, therefore, Chandrasekaran et al. started with the less complex small RNA subfraction. Wherein using the miR-206 target nanoswitch the scientists observed a progressive increase in miR-206 expression in small RNAs between undifferentiated samples and samples differentiated for 1-4 days.
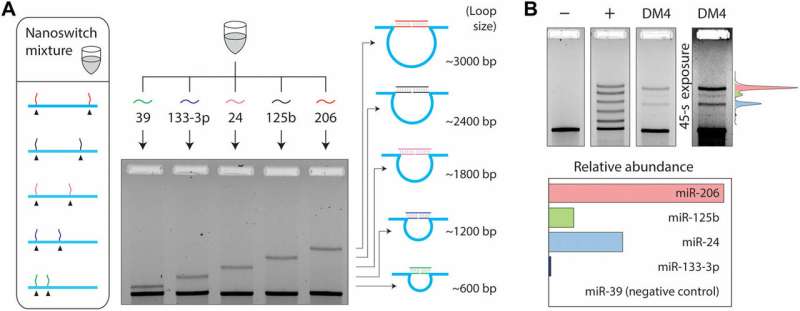
Since multiple mRNAs can also alter their expression during different cellular or disease stages, the phenomenon required additional capabilities of detection. To accomplish this in the experimental setup, Chandrasekaran et al. used the programmability of nanoswitches and developed a multiplexing system capable of detecting multiple miRNAs from the same sample. They placed the detector strands at desired locations of the DNA scaffold, resulting in loops of different sizes when bound to the target miRNA. The loop size of the nanoswitch thus determined gel migration, resulting in a unique band on the gel for accurate detection. For the experiment, the scientists chose four miRNAs present in muscle cells; miR-206, miR-125b, miR-24 and miR-133-3p and a negative control miRNA; miR-39 specific to the species Caenorhabditis elegans.
Within 50 ng of small RNAs, the scientists detected the four miRNAs at various expression levels, while confirming no detection with the negative control. The multiplexing strategy allowed the scientists to directly compare miRNA levels in a single sample, without labeling or amplification. In total, the work provided an additional step in the direction of expanding the throughput of the miRacles assay. The capability can be expanded to accommodate more miRNAs per switch as well.
In this way, Chandrasekaran et al. substantially moved forward from their preliminary proof-of-concept detection of synthetic DNA sequences; to establish, characterize and optimize a ready-to-use miRNA detection assay with biological extracts. The demonstrated work was a first example to use DNA nanoswitches to detect miRNAs from a true biological sample. While the performance of the miRNA assay was competitive compared to other commonly used techniques, the selectivity of 1 nucleotide seen in the present work was difficult to accomplish with existing methods. The sensitivity of miRacles also outperformed Northern blotting and microarrays. The assay could measure miRNAs without needing amplification, with simpler protocols and without the added error of extra sample processing. The protocol simply mixed the nanoswitches with the sample liquid for gel electrophoresis, to produce high quality results in the lab. The research work is potentially transferrable from biological to clinical samples to diagnose and monitor disease.
More importantly, the present work aligns with the broader concept of frugal science; a promising vision of cost effective science that has already produced low-cost solutions to blood centrifugation and water purification techniques in biomedical engineering. Chandrasekaran et al. aim to continue to contribute to the emerging trend in science, by disrupting the existing cost/performance relationship to provide broad scale access to simple miRNA detection methods with mix-and-read smart reagents.
More information: Arun Richard Chandrasekaran et al. Cellular microRNA detection with miRacles: microRNA- activated conditional looping of engineered switches, Science Advances (2019). DOI: 10.1126/sciadv.aau9443
Monya Baker. MicroRNA profiling: separating signal from noise, Nature Methods (2010). DOI: 10.1038/nmeth0910-687
M. Saad Bhamla et al. Hand-powered ultralow-cost paper centrifuge, Nature Biomedical Engineering (2017). DOI: 10.1038/s41551-016-0009
M. U. Sankar et al. Biopolymer-reinforced synthetic granular nanocomposites for affordable point-of-use water purification, Proceedings of the National Academy of Sciences (2013). DOI: 10.1073/pnas.1220222110
Journal information: Science Advances , Nature Biomedical Engineering , Nature Methods , Proceedings of the National Academy of Sciences
© 2019 Science X Network