Study provides new clues to improving chemotherapies
About half of all drugs, ranging from morphine to penicillin, come from compounds that are from—or have been derived from—nature. This includes many cancer drugs, which are toxic enough to kill cancer cells.
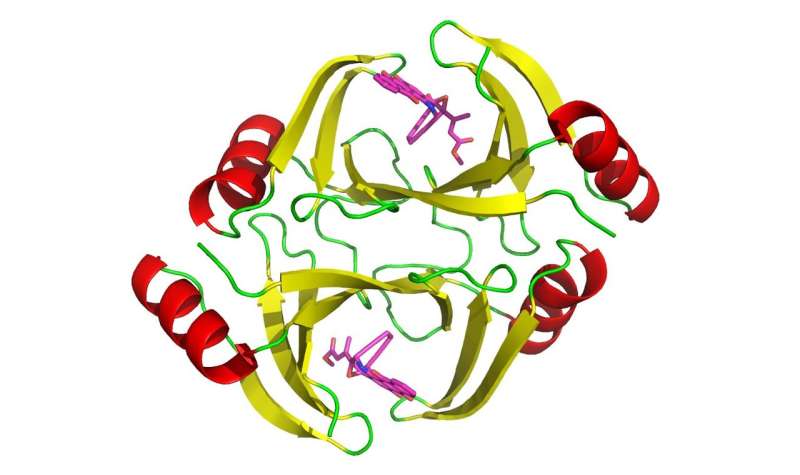
So how do the organisms that make these toxic substances protect themselves from the harmful effects? Scientists on the Florida campus of Scripps Research have uncovered a previously unknown mechanism—proteins that cells use to bind to a toxic substance and sequester it from the rest of the organism.
"Thanks to this discovery, we now know something about the mechanisms of resistance that's never been known before for the enediyne antitumor antibiotics," says study senior author Ben Shen, Ph.D., professor and co-chair of the Scripps Research Department of Chemistry.
The work has important implications for understanding how human cancer cells develop resistance to natural product-based chemotherapies. Furthermore, the microbiome may play a role in drug resistance. The study was published today in the journal Cell Chemical Biology.
"This mechanism could be clinically relevant for patients getting these drugs, so it's very important to study it further," says Shen.
Natural products—chemical compounds produced by living organisms—are considered one of the best sources of new drugs and drug leads. "They possess enormous structural and chemical diversity compared with molecules that are made in the lab," Shen says. Natural products may come from flowers, trees or marine organisms such as sponges. One of the most common sources, however, is soil-dwelling bacteria.
Shen's lab is focused on a class of natural products called enediynes. These compounds come from bacteria called actinomycetes, which are naturally found in the soil. Two enediyne products are already FDA approved as cancer drugs and are in wide use. But patients who take them often develop resistance. After a period of months or years, tumors can stop responding to the chemotherapy and begin growing again.
While how patients develop resistance to these drugs remains largely unknown, scientists have uncovered two mechanisms that bacteria use to protect themselves from enediynes. "Mechanisms of self-resistance in antibiotic producers serve as outstanding models to predict and combat future drug resistance in a clinical setting," says Shen.
In the new study, researchers report a third, previously undiscovered resistance mechanism. It involves three genes called tnmS1, tnmS2 and tnmS3, which encode proteins that allow bacteria to resist the effects of a type of enediynes called tiancimycins. Shen's lab is currently studying tiancimycins, which hold great promise for new cancer drugs. The proteins work by binding to tiancimycins and keeping them separate from the rest of the organism.
After discovering these genes in actinomycetes and how they work, the investigators studied how widespread these genes are in other microorganisms. They were surprised to find that in addition to actinomycetes, the genes were also present in several microorganisms commonly found in the human microbiota, the collection of microorganisms that naturally inhabit the human body.
"This raises a lot of questions that no one has ever asked before," Shen says. "I can rationalize why the producing organism would have these genes, because it needs to protect itself from its own metabolites. But why do other microorganisms need these resistance genes?" He notes that it may be possible for gut microbes to pass the products of these genes on to their host—humans—which could contribute to drug resistance.
"These findings raise the possibility that the human microbiota might impact the efficacy of enediyne-based drugs and should be taken into consideration when developing new chemotherapies," Shen says. "Future efforts to survey the human microbiome for resistance elements should be an important part of natural product-based drug discovery programs."
More information: Chin-Yuan Chang et al, Resistance to Enediyne Antitumor Antibiotics by Sequestration, Cell Chemical Biology (2018). DOI: 10.1016/j.chembiol.2018.05.012
Journal information: Cell Chemical Biology
Provided by The Scripps Research Institute