CRISPR-guided proximity labeling—dissecting gene transcription's proteomic choreography
More powerful than previous methods, a new approach for tagging and harvesting DNA-associated proteins from cells could open deeper insights into transcription control.
Gene transcription requires a complicated feat of molecular choreography, in which numerous proteins come together in the right place at the right time. Isolating, identifying, and studying those proteins is similarly complicated, and existing methods for doing so, while powerful, have limitations that restrict their utility.
The widely-used chromatin immunoprecipitation (ChIP) approach, for instance, relies on antibodies to fish proteins out of a cell. As such, it can only find proteins for which one has a high-quality, specific antibody and that one assumes may be present near a given gene, leaving out proteins that one might not expect.
In Nature Methods, a team led by Broad staff scientist Sam Myers, Broad Proteomics Platform director and institute scientist Steven Carr, and Broad core institute member Feng Zhang, revealed a new, unbiased approach for capturing proteins that help express a gene of interest, one that relies on a blunted version of the CRISPR-Cas9 genome editing system. Dubbed GLoPro (for genomic locus proteomics), the approach provides a more comprehensive view of which proteins manage a given gene's expression, knowledge that can help reveal insights into its function and its place in the cell's circuitry.
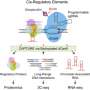
GLoPro relies on a catalytically inert (or "dead") form of the Cas9 protein (dCas9) fused to an engineered version of an enzyme called APEX2. Using guide RNAs, the fused protein—called CASPEX—homes in on a desired site in the genome in typical CRISPR fashion. Instead of cutting DNA, however, CASPEX labels, on command, any proteins or groups of proteins in its proximity with biotin. Researchers can then use this chemical tag to collect the labeled proteins for mass spectrometry-based proteomic analysis.
In their paper, the team showed that in cell lines they could label, capture, and analyze proteins bound to the promoters for the genes hTERT and MYC. Benchmarked against other protein isolation methods, GLoPro proved itself an effective and potentially generalizable way for collaring proteins near any location in the genome.
The team continues to explore the ways in which GLoPro could serve as a tool for gaining more comprehensive insights into the coordination of gene transcription.
More information: Samuel A. Myers et al. Discovery of proteins associated with a predefined genomic locus via dCas9–APEX-mediated proximity labeling, Nature Methods (2018). DOI: 10.1038/s41592-018-0007-1
Journal information: Nature Methods
Provided by Broad Institute of MIT and Harvard