Genetic screening tool identifies how the flu infiltrates cells
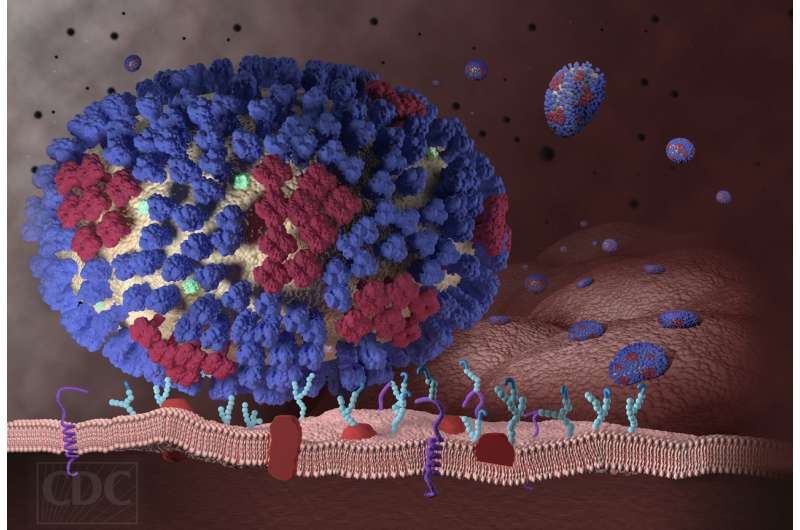
Researchers at the University of Chicago have developed a genetic screening tool that identified two key factors that allow the influenza virus to infect human lung cells. The technique uses new gene editing tools to create a library of modified cells, each missing a different gene, allowing scientists to see which changes impact their response to flu. This in turn could identify potential targets for antiviral drugs.
"Our current treatments for flu are limited. Vaccines have variable efficacy, and the virus has a propensity to mutate so that antiviral drugs don't work as well," said Julianna Han, a graduate student in microbiology at UChicago and lead author of the study, published April 10, 2018 in Cell Reports. "The next wave of antiviral treatments will be in part directed toward the host, so our work helps us get a better understanding of what proteins and pathways are utilized by the influenza virus."
Han and Balaji Manicassamy, PhD, assistant professor of microbiology at UChicago and senior author of the study, used CRISPR/Cas9 gene editing tools that allow scientists to selectively knock out, or turn off, specific genes. They created a library of modified human epithelial lung cells, the cells that line the airways and are the first to be infected by the flu virus. Each cell was missing a different gene, creating nearly 19,000 different genetic variations of the cell.
The researchers then exposed the cells to the H5N1 flu strain, a type of influenza A virus commonly known as the bird flu. If the virus was able to infect and kill one of the host cells, that means the gene and the proteins it produces didn't play a role in the virus' ability to replicate. If the cell survived, that means its modified genome somehow made it resistant to the virus—it was now missing a pathway that the virus relied on to replicate and do its dirty work.
After five rounds of exposing the cells to the H5N1 virus, Han and Manicassamy were left with a set of cells that were pretty resistant to the flu. When they examined what these hardy survivors had in common, two genes stood out. One, SLC35A1, encodes a protein that helps create a receptor for the flu on the surface of the cell. This makes sense—if you knock out SLC35A1, there's nothing for the virus to latch onto and infect the host.
The second gene, CIC, is more intriguing. It's a negative regulator of the innate immune system, meaning it helps shut down the cell's default immune response to foreign invaders. When CIC is turned off, other genes that produce antiviral and inflammatory responses are allowed to fire up and fight off the virus, which is why the test cells missing it were resistant to the flu.
But you couldn't just knock out a gene like CIC permanently as a means to fight the flu. The body needs mechanisms to shut off the immune system once an infection is gone. If not, it could go into overdrive and damage the body's own cells, which is what happens in autoimmune disorders. On the other hand, certain cancers can exploit a negative regulator like CIC to suppress immune responses while tumor cells run rampant.
"By turning off CIC, we now allow for expression of antiviral genes that are able to respond to the flu infection," Manicassamy said. "But it's also probably regulating inflammatory gene expression in autoimmunity and some cancers. It's a new twist that may be broadly applicable to other diseases besides the flu."
Once they identified the two key genes involved in H5N1 response, Han and Manicassamy exposed the cells to other pathogens to spot any more commonalities. CIC was also important for all strains of flu and several RNA viruses, or viruses containing RNA genomes. These included respiratory and non-respiratory viruses, highlighting CICs broad effect.
The new genetic screening process could be used to find other host genes relevant to different viruses that infect epithelial lung cells. Then, researchers could look for any currently available drugs that target those pathways and see if they can be repurposed as antiviral treatments.
Ultimately, Han is working to understand exactly how CIC regulates other antiviral and inflammatory genes. How does the host turn CIC on or off? And what happens when it malfunctions?
"It's either through mutations in CIC itself or through aberrant regulation," she said. "If we can further pinpoint what is involved, we can try to figure out how CIC is being misregulated in cancer and autoimmune diseases."
More information: Julianna Han et al, Genome-wide CRISPR/Cas9 Screen Identifies Host Factors Essential for Influenza Virus Replication, Cell Reports (2018). DOI: 10.1016/j.celrep.2018.03.045
Journal information: Cell Reports
Provided by University of Chicago Medical Center