Double-bridged peptides bind any disease target
Peptides are short chains of amino acids that can bind to proteins and change their function. They show high binding affinity, low toxicity, and are easy to synthesize, all of which makes peptides ideal for use in drug development, and many naturally occurring peptides such as insulin, oxytocin, somatostatin and the antibiotics vancomycin or polymyxin B, are widely used.
However, using peptides as drugs faces two challenges. First, binding affinity: Good binding requires difficult peptide architectures and amino acid sequences that are perfectly complementary in shape and charge to the surface of their target proteins. Second, peptide stability: peptides can potentially be cut by enzymes (proteases) into smaller, useless fragments or even single amino acids.
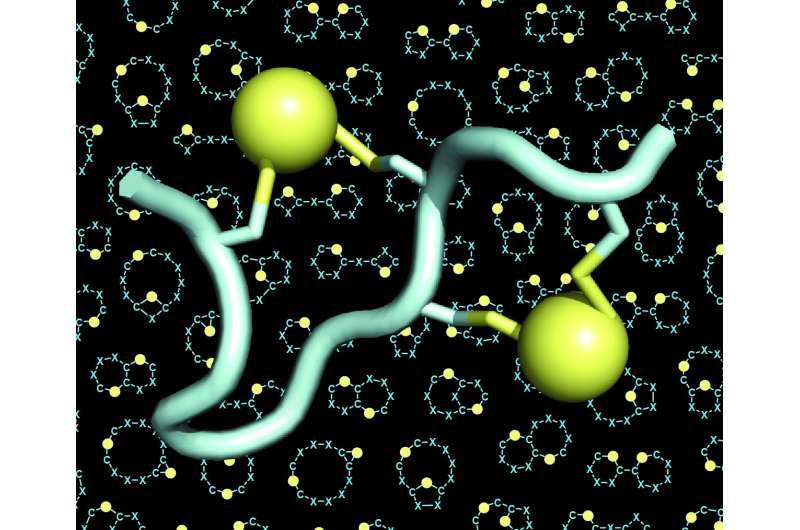
The lab of Christian Heinis at EPFL has now addressed both of these challenges by developing the new peptide format that they call "double-bridged peptides." These are chains of 10 to 15 amino acids, four of which are chemically connected by two bridges. Each bridge links a pair of two cysteine amino acids—four in total.
Because the four cysteines can be placed in many different configurations along the sequence of amino acids, the double-bridging strategy allowed the researchers to generate an enormously large number of structurally diverse peptide architectures. They expanded the diversity even further by using different chemical reagents that increased the number of bridges. The scientists also found that the four cysteines could be bridged by two linkers in three different ways, giving rise to three different architectures from each single peptide sequence.
Using this strategy, the researchers produced an enormous diversity of peptide structures, all with different "skeleton" structures. In addition, the chemists systematically changed the amino acids between the cysteines and generated libraries of billions of different double-bridged peptides. After screening the libraries, the researchers were able to isolate high-affinity binders to important protein targets. One of these targets was kallikrein, a plasma protein that is associated with hereditary angioedema, a rare swelling disorder. Another was interleukin-17, a cytokine protein implicated in several inflammatory disorders such as rheumatoid arthritis and psoriasis. Using the double-bridge approach for both these protein targets, the scientists developed peptides that could bind them efficiently at nanomolar concentrations. In the case of kallikrein, the peptides could bind it for more than an hour before dissociating.
Stability was also a highly attractive feature of the new peptide format. The double-bridged peptides are hardly degraded by proteases in the blood, which is a great advantage because it keeps the peptides from being eliminated too quickly, thus extending their therapeutic effects.Based on the results, Heinis's lab is now applying the peptide format to many other disease targets. They have already developed new, even larger double-bridged peptide libraries and screened them against a range of disease-relevant targets. One of these peptides is already undergoing pre-clinical evaluation.
More information: Sangram S. Kale et al, Cyclization of peptides with two chemical bridges affords large scaffold diversities, Nature Chemistry (2018). DOI: 10.1038/s41557-018-0042-7
Journal information: Nature Chemistry
Provided by Ecole Polytechnique Federale de Lausanne





















