Scientists discover a molecular timer based on stalling ribosomes
A molecular biologist from Lomonosov Moscow State University and international colleagues discovered a special mechanism of protein synthesis regulation that they called a "molecular timer." It controls the number of protein molecules produced by a cell and prevents the generation of extra molecules. When activated with drugs, such a timer may help efficiently combat cancerous tumors. The study was reported in Nature.
Genetic information coded in DNA is copied into the molecules of messenger RNA (mRNA), after which proteins are synthesized. Each amino acid of a protein corresponds to three nucleotides (one triplet) in mRNK. An organelle responsible for protein synthesis is called a ribosome. It is a molecular machine that consists of subparticles, one larger and one smaller (each comprised of ribosome RNA and numerous proteins). A signal to end protein synthesis is given by three stop codons. In the majority of living bodies these are UGA, UAA, and UAG (U—uracil, A—adenine, G—guanine). A molecule of mRNA is always longer than the reading frame—an element that contains genetic information about a protein. The elements that do not code any protein-related information are called untranslated (or non-coding) regions. In mammals, they are often much larger than reading frames and can contain thousands of nucleotides. Given that mRNA synthesis is an energy-consuming process, scientists have been wondering about the purpose of these huge, untranslated regions.
Today we know that non-coding regions regulate the life cycle of mRNA. Unlike DNA, mRNA molecules are destroyed in the cell after some time. Some of them live quite long after synthesis, and others break down in a couple of minutes. Untranslated regions of mRNA often play a big role in this process.
Another important function of such regions is the regulation of protein synthesis. There are many examples of non-coding mRNA regions being bound with regulatory proteins (or short RNA) that either suppress or stimulate protein synthesis, thus allowing the cell to turn protein synthesis on a certain mRNA molecule on or off. It is an important function, as depending on the environmental conditions, a cell needs a certain set of proteins. Any abnormalities in synthesis regulation may cause uncontrollable protein generation. For example, deregulation of proteins in charge of cell division may lead to endless growth of new cells, which is characteristic for cancer.
Having studied the regulation of mRNA translation of Amd1 enzyme, a key element in the biological synthesis of polyamines (polymers containing amine groups), a team of scientists from University College Cork, Harvard Medical School, Lomonosov Moscow State University and the Department of Human Genetics at the University of Utah discovered a new mechanism of protein synthesis regulation.
"We've known for a long time that it is hard for ribosomes to synthesize the sequences of certain triplets," explains Dmitry Andreev, a senior research associate of Belozersky Science and Research Institute of Physical and Chemical Biology, MSU. "Therefore, when encountering such a sequence, a ribosome may get stalled on an mRNA molecule. This causes all following ribosomes to stop as well, as they cannot jump over their stalled buddy, and protein synthesis is suspended. In the course of our studies of Amd1 regulation, we discovered a stalling signal. We were interested why it was located after the stop codon of the Amd1 reading frame. What was it doing there?"
The translation mechanism should prevent any errors in protein synthesis as the wrong proteins can cause harm to the cell. However, its precision is limited. A ribosome that reaches a stop codon may read it as an amino acid by mistake and move further, synthesizing a longer protein. The probability of this happening is quite low. According to the scientists, it amounts to a fraction of a percent. Still, the reading of the stop codon was a key to understanding the new Amd1 mechanism regulation.
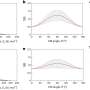
The mechanism was discovered using ribosome profiling, a method based on mass identification of the nucleotide (genetic) sequence of mRNA fragments that react with ribosomes. The methodology allows for studying protein synthesis in a cell on the full genome level. Later on, the mechanisms were studied with the use of traditional reporter constructions—clusters made of a gene in question and a reporter gene that shows the level of gene expression in a cell. The reporter genes in this case were luciferase (an enzyme that triggers a reaction causing light emission) and green fluorescent protein (GFP).
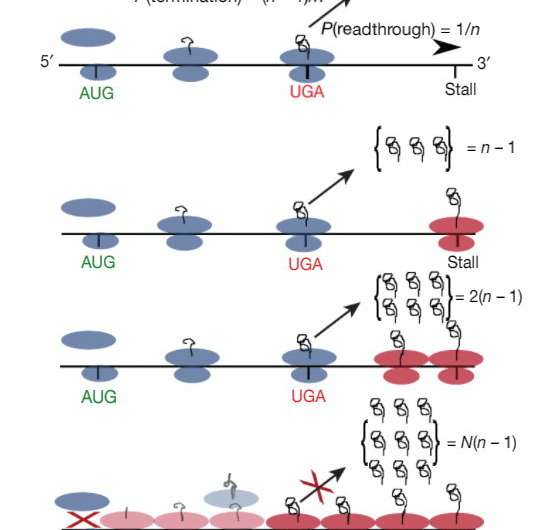
The team found that approximately one in 60 ribosomes moves through the stop codon of Amd1, but after some time, it stalls on the "wrong" sequence. This is not bad in isolation, but the next ribosome that moves through the stop codon stalls behind it. The ribosomes queue until the line reaches the stop codon. As soon as it happens, the production of protein stops.
"We called this regulation mechanism a molecular timer," says Andreev. "Such an mRNA molecule has a built-in mechanism that regulates the number of synthesized protein molecules very precisely. Even if ribosomes frequent this mRNA, the reading of the stop codon and further stalling (the molecular timer) will switch protein synthesis off after a number of cycles."
Amd1 has oncogenic properties. It has already been demonstrated that its excessive production in a cell may lead to the growth of very aggressive and metastasizing tumors. If we learn to turn on the timer that regulates its synthesis (for example, to potentiate the reading of the stop codon) with drugs, this may become a new therapeutic approach. Some medicinal drugs based on the activating of stop codon reading are already being developed, and one of them has been approved in Europe for treating Duchenne's dystrophy.
More information: Martina M. Yordanova et al, AMD1 mRNA employs ribosome stalling as a mechanism for molecular memory formation, Nature (2018). DOI: 10.1038/nature25174
Journal information: Nature
Provided by Lomonosov Moscow State University