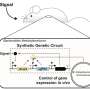
Red, green, and blue light can be used to control gene expression in engineered E. coli

MIT researchers have engineered bacteria with "multicolor vision"—E. coli that recognize red, green, or blue (RGB) light and, in response to each color, express different genes that perform different biological functions.
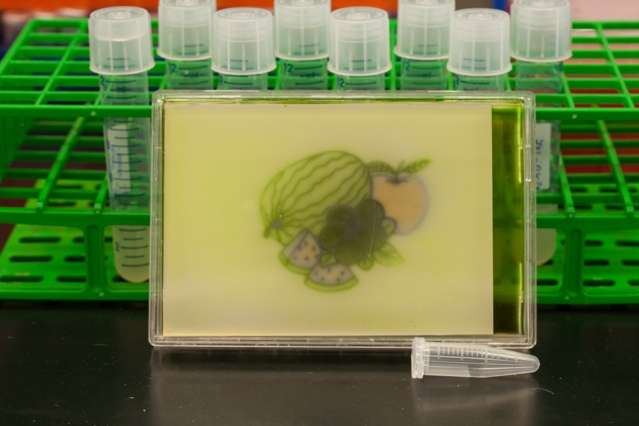
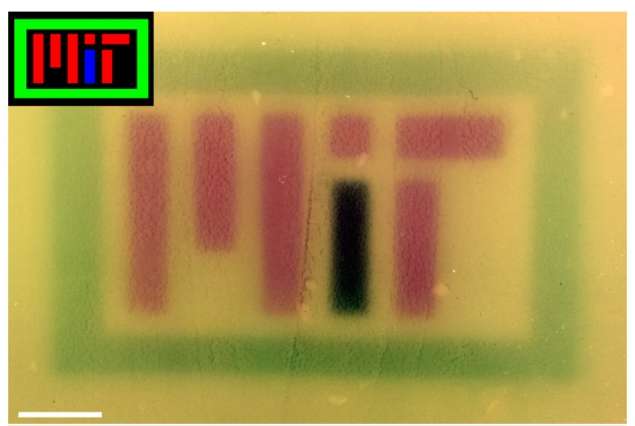
To showcase the technology, the researchers produced several colored images on culture plates—one of which spells out "MIT"—by using RGB lights to control the pigment produced by the bacteria. Outside of the lab, the technology could also prove useful for commercial, pharmaceutical, and other applications.
The E. coli is programmed with a protein- and enzyme-based system, analogous to a computer chip, with several different modules to process the light input and produce a biological output. In computing terms, a "sensor array" first becomes activated in the presence of either red, green, or blue light, and a "circuit" processes the signal. Then, a "resource allocator" connects the processed information to "actuators" that implement the corresponding biological function.
Think of the new E. coli as microbial marionettes, with colored light instead of puppet strings making the bacteria act in a certain way, says MIT professor of biological engineering Chris Voigt, co-author of a paper in Nature describing the technology. "Using different colors, we can control different genes that are being expressed," he says.
The paper's co-authors are former postdocs Jesus Fernandez-Rodriguez, Felix Moser, and Miryoung Song.
Synthetic-biology innovation comes together
In 2005, Voigt, who co-directs the Synthetic Biology Center at MIT, and other researchers pioneered a "bacterial camera" by programming a light sensor into a strain of E. coli, along with a gene that produced black pigment. When light shone through a stencil onto a bacteria-coated plate, the microbes formed black-and-white images. At the time, this feat required only four genes and three promoters—regions of DNA that initiate gene transcription—to get the job done.
New synthetic biology tools, such as the genome-editing system CRISPR, have cropped up since then, opening broader possibilities to researchers. In contrast to the 2005 system, the new RGB system—the first to use three colors—consists of 18 genes and 14 promoters, among other parts, as well as 46,000 base pairs of DNA.
But with greater complexity come greater challenges. Because the researchers were dealing with a sensor array that could detect three separate colors, for instance, they had to include in the microbial program a protein that prevents gene transcription of the two unused sensors.
In computing terms, this is called a "NOT gate," a circuit that produces an output signal—in this case, gene repression—only when there is not a signal on its input. With bacteria under a red light, for instance, the NOT gate would unleash that gene-repressing protein on the green and blue sensors, turning them off.
About five years ago, Voigt led a team that engineered microbes to respond to red and green light. Adding a third sensor was a major challenge of the new research. "Inside the cell, all the new protein sensors you add interfere with each other, because it's all molecules bumping around the cell, and they all require keeping the cell alive and happy. With every additional sensor you add, that gets exponentially harder," he says.
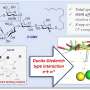
In that regard, Voigt adds, the system's resource allocator, a new feature, also acts as a circuit breaker, shutting down the sensors if all three turn on at once, overloading the cell.
From a genetic engineering perspective, the four-subsystem configuration was "the biggest impact of this work," Voigt says. Each subsystem—the sensor array, circuits, resource actuators, and actuators—was designed, built, and optimized in isolation before being assembled into a final structure. This simplified, modular process could pave the way for more complex biological programming in the future, according to the researchers.
Generally speaking, Voigt sees the new system as a culmination of a decade of synthetic-biology innovations. "It's a representation of where we are currently, and all the pieces that needed to come together over the last decade to create systems of this scale and complexity," he says.
Making "disco bacteria"
To make the new color images, the researchers programmed bacteria to produce the same pigment as the red, green, or blue light shone upon them. In an incubator, the researchers coated a petri dish with bacteria that are genetically identical. "You can think of it like undeveloped film, where you have the petri dish with bacteria on it," Voigt says, "and the camera is the incubator."
At the top of the incubator is a hole, where a stenciled image is projected onto the plate. Over time, the bacteria grow, producing an enzyme that produces a pigment corresponding to whichever RBG color they're illuminated by. In addition to the MIT logo, the researchers produced images of various patterns, multicolored fruit, and the video game character Super Mario.
The engineered bacteria could also be used to rapidly start and stop the chemical reactions of microbes in industrial fermentation processes, which are used to make pharmaceuticals and other products. Today, controlling such chemical reactions requires dumping different chemical additives into large fermenting vats, which is time-consuming and inefficient.
In their paper, the researchers demonstrated this "chemicals on-demand" concept on a small scale. Using CRISPR gene-editing tools, they modified three genes that produce acetate—a sometimes-unwanted byproduct of various bioprocesses—to produce less of the chemical in response to RGB lights.
"Individually, and in combination with one another, the different colors of light reduce acetate production without sacrificing biomass accumulation," the researchers wrote in their paper.
Voigt has coined an amusing name for these industrial microbes. "I refer to them as 'disco bacteria,'" he says, "because different colored lights are flashing inside the fermenter and controlling the cells."
A future application, Voigt adds, could be in controlling cells to form various materials and structures. Researchers, including some at MIT, have started programming cells to assemble into living materials that one day could be used to design solar cells, self-healing materials, or diagnostic sensors.
"It's amazing when you look at the world and see all the different materials," Voigt says. "Things like cellulose, silk proteins, metals, nanowires, and living materials like organs—all these different things in nature we get from cells growing into different patterns. You can imagine using different colors of light to tell the cells how they should be growing as part of building that material."
More information: Jesus Fernandez-Rodriguez et al. Engineering RGB color vision into Escherichia coli, Nature Chemical Biology (2017). DOI: 10.1038/nchembio.2390
Journal information: Nature Chemical Biology
Provided by Massachusetts Institute of Technology
This story is republished courtesy of MIT News (web.mit.edu/newsoffice/), a popular site that covers news about MIT research, innovation and teaching.