First large-scale interactome map of the largest membrane receptors group in humans
A team of the Institute of Neurosciences of the University of Barcelona has participated in the design of the first large-scale interaction map of G-protein-coupled receptors in humans, the largest group of membrane proteins that control essential functions of cells (metabolism, proliferation, differentiation, etc.).
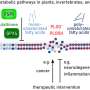
The new interaction map or GPCRs interactome, the largest described so far in this group of proteins, will be a step forward to discover the origins of some neurological pathologies (Parkinson, Alzheimer, schizophrenia, epilepsy and some neurogliomas). Knowing with details about the connection of these proteins will also help to identify new therapeutic targets, design future drugs and understand the negative effects associated with current medicines.
This research, published in Molecular System Biology, and led by Igor Stagljar, from the University of Toronto (Canada), has the participation of the researchers Francisco Ciruela, Jorge Gandía and Xavier Morató, from the Faculty of Medicine and Health Sciences of the University of Barcelona, the Institute of Neurosciences of the University of Barcelona, and the Bellvitge Biomedical Research Institute (IDIBELL), among other experts.
Interactome: how proteins interact in living beings
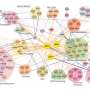
Any protein acts by itself, therefore it is essential to study interactions between proteins to understand the complexity of the network of cell and physiological functions in living beings. Knowing the topology and connection patterns between proteins gives an essential perspective to discover the systemic biology of living beings. The idea of interactome –the protein function interaction network- was defined for the first time in 1999 (Sánchez et al., Nucleics Acids Research), and in 2000 the first results of interactome or protein-protein interactions were published (Schwikowski et al, Nature Biotechnology).
G-protein-coupled receptors (GPCRs) are a family of hundreds of biological membrane proteins involved in cell signal transduction. These structures are fully absorbed in cell membranes, which hardens their study and manipulation in experimental studies.
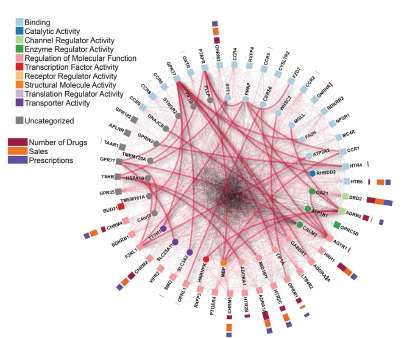
"This new study shows the first large-scale map of interactome for GPRCs family in humans. In total, the interactome describes new 987 protein-protein interactions in which GPCRs take part and which occur within 686 proteins, 299 out of which are membrane integral proteins" says Professor Francisco Ciruela.
MYTH approach: inflection point in experimental methodology
Traditionally, in order to determine interactomes in an experimental way, there were two methodology approaches: the conventional yeast two-hybrid (YTH) system, and affinity purification-linked to mass spectrometry (AP-MS). However, in some cases the applied methodology alters the features of the cell environment and therefore, the proteins' and interactome's too. The yeast two-hybrid technique (YTH), an innovative protocol applied in the new study, has solved this severe experimental limitation.
"Thanks to the YTH protocol –says Ciruela- which is a modification of the classic yeast two-hybrid system (YTH), we could study membrane integral proteins such as the GPCRs, in their membrane cell environment. Apart from other obtained results, we could define the interactome of 48 different GPCRs with the YTH technique".
Interactome networks and functional interactions between proteins
Interactomes and biological networks are advanced elements in basic research to identify possible targets of pharmacological interest. In order to work on these interactome networks of high-fidelity proteins, it is necessary to establish the interaction and functional association between proteins and complementary procedures. In the new study, the interactome map defined with the YTH system has passed a second validation round with the YTH system, which enables defining a precise list of interaction partners for each receptor. Then, an important number of these interactions are confirmed in human cell cultures. Therefore, 40 interactions described by YTH were validated.
Exploring the therapeutic potential to treat Parkinson
In one of the last stages of the study, two of the described connections linked to Parkinson's disease were functionally validated: the one in serotonin 5-HT4d with receptors GPRIN2 and GPR37 –a work from the team led by Ralf Jockers, from the University Paris-Descartes (France)- and the one of the A2A adenosine receptor and receptor GPR37, a distinguished scientific contribution by the team of the UB and IDIBELL in the international study.
The experts of UB-IDIBELL characterized the molecular interaction of the A2A adenosine receptor with the orphan receptor GPR37 –a molecule associated with the death of dopaminergic neurons in Parkinson's disease- an animal model of psychomotor behaviour (in particular, in a catalepsy model induced by the blocking of dopaminergic transmission).
According to the conclusions, animal models that don't express the GPR37 receptor undergo a higher efficacy from antiparkinson drugs (such as antagonists of the A2A adenosine receptor) in their pro-dopaminergic activity.
The research lines unfolded by the team of the UB-IDIBELL aim to discover the function of GPR37, its role in the origin of Parkinson's disease and its potential as a new therapeutic target to treat this severe neurological disease.
More information: Kate Sokolina et al. Systematic protein–protein interaction mapping for clinically relevant human GPCRs, Molecular Systems Biology (2017). DOI: 10.15252/msb.20167430
Journal information: Nature Biotechnology , Molecular Systems Biology
Provided by University of Barcelona