New big data approach predicts drug toxicity in humans
Researchers can now predict the odds of experimental drugs succeeding in clinical trials, thanks to a new data-driven approach developed by Weill Cornell Medicine scientists. The method detects toxic side effects that may disqualify drugs from human use, giving drug developers an early warning before initiating clinical trials, according to a new study published Sept. 15 in Cell Chemical Biology.
The approach upends conventional wisdom about the criteria on which to evaluate new drugs for their safety. Rather than exclusively examining molecular structure to determine viability, this new computational method combines a host of structural features and features related to how the drug binds to molecules in the body.
"We looked more broadly at drug molecule features that drug developers thought were unimportant in predicting drug safety in the past. Then we let the data speak for itself," said author Dr. Olivier Elemento, an associate professor of physiology and biophysics and of computational genomics in computational biomedicine, associate director of the HRH Prince Alwaleed Bin Talal Bin Abdulaziz Al-Saud Institute for Computational Biomedicine, and head of the computational biology group at the Caryl and Israel Englander Institute for Precision Medicine.
The method, known as PrOCTOR, was inspired by an approach that baseball statisticians adopted to better predict which players would be successful offensively. Instead of relying on collective wisdom from baseball insiders, statisticians decided to use an objective numbers analysis to measure in-game productivity, a strategy known as "Moneyball."
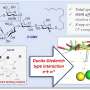
Similarly, researchers developed a computational method that analyzes data from 48 different features of a drug—from molecular weight to details about its target—to determine whether it would be safe for clinical use. Using a form of artificial intelligence called machine learning, the investigators trained PrOCTOR on hundreds of U.S. Food and Drug Administration-approved drugs and drugs that failed clinical trials due to toxicity problems.
Based upon this information, the investigators created "PrOCTOR scores" that could help distinguish drugs approved by the FDA from those that failed for toxicity. They tested PrOCTOR on hundreds of additional drugs approved in Europe and Japan and using side-effect data on approved drugs collected by the FDA. PrOCTOR was able to accurately recognize toxic side effects that were a consequence of a drug's chemical features and its target. Records revealing that many of these drugs had failed clinical trials supported PrOCTOR's accuracy.
"We were able to find several features that led to a very predictive model," Dr. Elemento said. "Hopefully this approach could be used to determine whether it's worth pursuing a drug prior to starting human trials."
He added that the method could also be utilized for post-approval surveillance of drugs that are currently approved by the FDA but may still be toxic. For example, PrOCTOR predicted that an FDA-approved diabetes drug was toxic, and upon further investigation, Dr. Elemento and his team discovered that it had been withdrawn from European markets.
But when it comes to toxicity, first author and doctoral candidate Kaitlyn Gayvert said context is vital. "A good example of this is chemotherapy," said Gayvert, who was named as one of Forbes' 30 Under 30 last year for her work on the project. "When treating advanced cancer, there is a higher bar for the types of side effects that doctors are going to allow."
She said this approach could improve the drug discovery pipeline, save money and save lives—but only if more data on toxicity results become available. After all, only 50 percent of clinical trial results are fully reported, Dr. Elemento said, adding that, "if we don't have data, we can't build these models."
More information: Kaitlyn M. Gayvert et al. A Data-Driven Approach to Predicting Successes and Failures of Clinical Trials, Cell Chemical Biology (2016). DOI: 10.1016/j.chembiol.2016.07.023
Journal information: Cell Chemical Biology
Provided by Cornell University