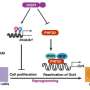
Scientists develop pioneering method to define stages of stem cell reprogramming
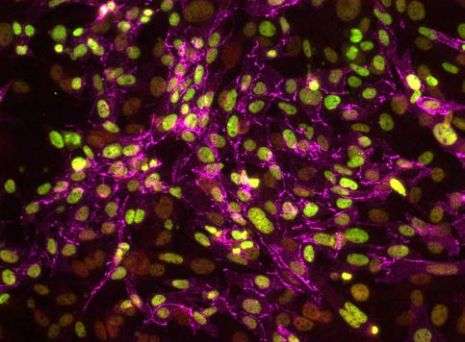
In a groundbreaking study that provides scientists with a critical new understanding of stem cell development and its role in disease, UCLA researchers at the Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research led by Dr. Kathrin Plath, professor of biological chemistry, have established a first-of-its-kind methodology that defines the unique stages by which specialized cells are reprogrammed into stem cells that resemble those found in the embryo.
The study was published online ahead of print in the journal Cell.
Induced pluripotent stem cells (known as iPSCs) are similar to human embryonic stem cells in that both cell types have the unique ability to self-renew and have the flexibility to become any cell in the human body. iPSC cells, however, are generated by reprogramming skin or blood cells and do not require an embryo.
Reprogramming is a long process (about one to two weeks) and largely inefficient, with typically less than one percent of the primary skin or blood cells successfully completing the journey to becoming an iPSC. The exact stages a cell goes through during the reprogramming process are also not well understood. This knowledge is important, as iPSCs hold great promise in the field of regenerative medicine, as they can provide a single source of patient-specific cells to replace those lost to injury or disease. They can also be used to create novel disease models from which new drugs and therapies can be developed.
"This research has broad impact, because by deepening our understanding of cell reprogramming we have the potential to improve disease modeling and the generation of better sources of patient-specific specialized cells suitable for replacement therapy," said Plath. "This can ultimately benefit patients with new and better treatments for a wide range of diseases.
Drs. Vincent Pasque and Jason Tchieu, postdoctoral fellows in the lab of Dr. Plath and co-first authors of the study, developed a roadmap of the reprogramming process using detailed time-course analyses. They induced the reprogramming of skin cells into iPSC, then observed and analyzed on a daily basis or every other day the process of transformation at the single-cell level. The data were collected and recorded over a period of up to two weeks.
Plath's team found that the changes that happen in cells during reprogramming occur in a sequential stage-by-stage manner, and that importantly, the stages were the same across all the different reprogramming systems and different cell types analyzed.
"The exact stage of reprogramming of any cell can now be determined," said Pasque. "This study signals a big change in thinking, because it provides simple and efficient tools for scientists to study stem cell creation in a stage-by-stage manner. Most studies to date ignore the stages of reprogramming, but we can now seek to better understand the entire process on both a macro and micro level."
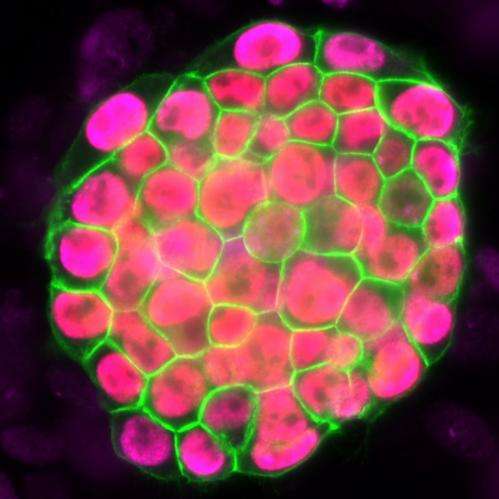
Plath's team further discovered that the stages of reprogramming to iPSC are different from what was expected. They found that it is not simply the reversed sequence of stages of embryo development. Some steps are reversed in the expected order; others do not actually happen in the exact reverse order and resist a change until late during reprogramming to iPSCs.
"This reflects how cells do not like to change from one specialized cell type to another and resist a change in cell identity," said Pasque. "Resistance to reprogramming also helps to explain why reprogramming takes place only in a very small proportion of the starting cells."
With these findings, Plath's lab plans future studies to actively isolate specific cell types during specific stages of reprogramming. They also hope the research will encourage further investigation into the characteristics of iPSC development.
More information: "X Chromosome Reactivation Dynamics Reveal Stages of Reprogramming to Pluripotency," Cell, Volume 159, Issue 7, 18 December 2014, Pages 1681-1697, ISSN 0092-8674, dx.doi.org/10.1016/j.cell.2014.11.040.
Journal information: Cell
Provided by University of California, Los Angeles