Seeing the messages microbes send: Novel chemical imaging instrument shows how bacteria support nearby colonies

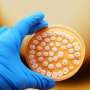
(Phys.org) —With a novel technique that noninvasively analyzes microbes, scientists at Pacific Northwest National Laboratory profiled, for the first time, the chemicals that a cyanobacterium makes available to others. Over 4 days, Synechococcus sp. PCC 7002 steadily secretes two molecules that could be used as resources by other bacteria that are nearby. The technique that chemically profiles the microbial communities in both space and time is Nanospray Desorption Ionization Electrospray Mass Spectrometry, or nano-DESI. This instrument was built by Dr. Julia Laskin and her team at Pacific Northwest National Laboratory. This research graced the cover of Analyst.
"This is a tool that will help microbiologists identify molecules that promote or inhibit growth of microbial communities," said Lab Fellow Laskin. "It also gives us much better control for studying interactions between microbial communities."
Understanding microbial ecology—how bacteria, algae and other microbes influence each other—could provide basic answers needed to advance sustainable energy. For example, Synechococcus sp. PCC 7002 uses carbon dioxide and sunlight to produce sugars that fuel the colony. Knowing how to best grow and modify these bacteria to mass-produce fuels could increase our nation's energy independence. Here, nano-DESI provides key data for sustainable energy, but the opportunities stretch much farther.
"Any place where there are microbes and you have a format where nano-DESI could be applied, you can study that ecology," said Dr. Allan Konopka, a biologist and Lab Fellow at PNNL who worked on the study. "This opens doors to a host of applications, such as understanding how bacteria associated with plant roots affect a plant."
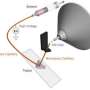
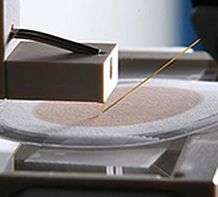
One of the advantages of a multidisciplinary national laboratory environment, where teams are constantly solving different problems for different clients, is the opportunity for synergistic connections. Laskin and her team began developing nano-DESI in 2009. The technique uses two thin glass capillaries. The input capillary produces a small drop of liquid that touches the microbial colony. The output capillary draws up liquid off the drop, which now contains chemicals produced by the colony. The sample-laden liquid is sent to a mass spectrometer for analysis.
"The technique allows rapid, repeated analysis of microbial colonies without destroying the colony as conventional methods would," said Dr. Ingela Lanekoff, an analytical chemist at PNNL who spent a year obtaining and analyzing the data. "It is really exciting to be seeing chemical snapshots of living microbial communities and their surrounding environment."
Konopka realized the potential of nano-DESI to study the molecules a microbe emits. The team began with Synechococcus sp. PCC 7002. The microbe influences nearby Shewanella colonies, which thrive, but only after the Synechococcus have been growing for several days. The team wanted to know how the Synechococcus was promoting the Shewanella.
The team placed the Synechococcus in a single line on high-salt nutrient plates. This approach enabled characterization of the spatial distribution of the communicating chemicals using nano-DESI. "It isn't enough to detect and identify molecules," said Laskin. "We need spatial profiling."
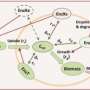
The team detected and identified metabolites and lipids on the colony and nearby. Using the nondestructive and highly sensitive nano-DESI, the team also identified several glycolipids that had not been detected by more conventional methods.
The researchers continually sampled the colonies over several days and measured the spread or chemical gradient of sucrose and glucosylglycerol, which protects the colony from the surrounding salt. "After a few days, glucosylglycerol was spread all over the agar plate," said Lanekoff.
They found that the distribution of both molecules on the surrounding medium changed as the colony aged.
"This is just a first step," said Konopka. "This technology allows us to characterize the most relevant environment for a microbe, which extends just a millimeter or less, and gives us the chance to look at microbial interactions mechanistically in a very precise way."
Nano-DESI will continue to aid biologists and others in characterizing molecular-level interactions between bacterial colonies, providing information that may be of use to biofuel production, disease treatment, and water and soil cleanup.
More information: Lanekoff, I. et al. 2013. Spatially Resolved Analysis of Glycolipids and Metabolites in Living Synechococcus Sp. PCC 7002 Using Nanospray Desorption Electrospray Ionization. Analyst. DOI: 10.1039/C3AN36716A
Journal information: Analyst
Provided by Pacific Northwest National Laboratory