Modeling microbes to manage carbon dioxide
(PhysOrg.com) -- In the past decade, microbiologists began realizing that communities of microbes process energy and materials, which affects their environments. To understand how microbial communities function in a natural ecosystem, Pacific Northwest National Laboratory scientists developed a novel kinetic model that represents microbial community dynamics in soil pores.
"Modeling the dynamics of soil bacterial communities is extremely challenging because it involves disparate physical, chemical, and biological components," said Dr. Allan Konopka, director of PNNL's Microbial Communities Initiative, or MCI. The Laboratory-funded MCI began in 2009 to advance the understanding of how microbial communities function in a natural ecosystem.
"In this study, we've developed a framework for a unified model that can incorporate a broader range of physical, chemical, and biological heterogeneities," Konopka continued. "This model will let us evaluate the consequences of different strategies that microbes might use for degrading organic matter, such as cellulose in soil, and analyze how three-dimensional pore structure impacts biological activity."
The results of this modeling work were published online by Microbial Ecology.
Cellulose is the most abundant naturally occurring material on Earth and is a primary structural component of plants. The microbial breakdown of cellulose and related byproducts is a key process in the global carbon cycle. Each year, it is estimated that more than 100 billion tons of cellulose are synthesized by plants and subsequently degraded by microorganisms that, in turn, produce carbon dioxide via respiration.
"Within the MCI, we are investigating the biogeochemical processes at the microscale because these have macroscopic impacts on the Department of Energy's carbon management mission," Konopka said. "Key to this is to study microbial communities—both in the natural environment and under better-controlled conditions in the lab. And, a key factor for lab-based studies is to represent the spatial heterogeneity found in soil aggregates and then to have simulation models that can incorporate the rules we've learned to predict behavior in the field."
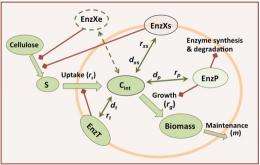
The model incorporates certain biological and physical properties of bacterial communities. Dr. Haluk Resat, one of the model's developers, explained: "The unique feature of our model is that it factors in a range of possible microsite conditions while accounting for heterogeneity in a three dimensional system and treating the metabolism of microbial cells individually based on their own local environmental conditions and history."
This model allowed the scientists to explicitly simulate microbial diversity, as well as to model cell physiology and how it is regulated. They simulated microbial community dynamics at two spatial scales: micropores, resembling 6- to 20-micrometer portions of the soil physical structure, and 111-micrometer soil aggregates with a random pore structure. The scientists observed an emergent dynamic behavior in bacterial communities consisting of organisms with complementary cellulose degradation mechanisms, where coexistence of different degradation functionalities led to more efficient cellulose utilization kinetics and more stable communities.
The scientists recognize the current model is only an initial step toward the creation of a comprehensive multiscale model for microbial dynamics that combines diversity in cellular metabolism and physiology with the physical and chemical constraints and heterogeneities occurring in the soil environment. As such, Resat and his colleagues are working on scaling up the developed model to "aggregate of aggregates" to study microbial kinetics in a representative bulk sample.
"Even at the present level of development, the simulation results demonstrate that this model captures some essential kinetic properties of bacteria and can be used as a powerful tool to generate experimentally testable hypotheses," Konopka said.
More information: Resat H, V Bailey, LA McCue, and A Konopka. In press. "Modeling Microbial Dynamics in Heterogeneous Environments: Growth on Soil Carbon Sources." Microbial Ecology (Online First), published online December 23, 2011.
Provided by Pacific Northwest National Laboratory