This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers reprogram bacterial gene activity with red light
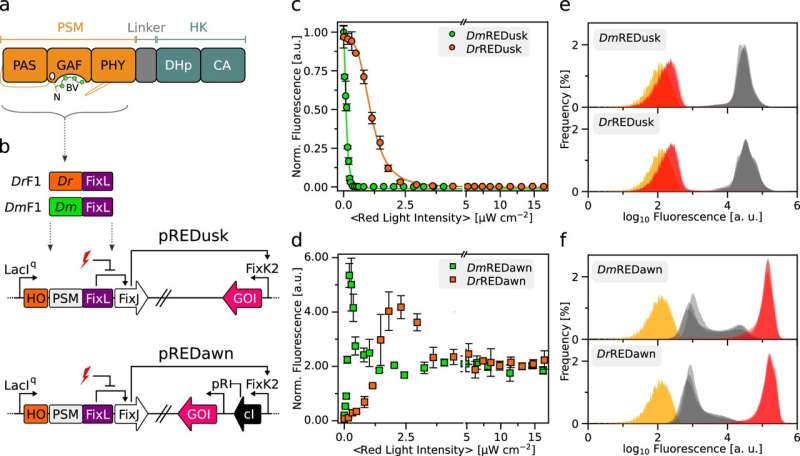
Researchers at the University of Bayreuth have changed the sensitivity of bacterial systems for controlling gene activity to red light and reprogrammed their molecular response to the light stimulus. The results, published in Nature Communications, open up exciting possibilities in the biotechnological application of bacteria.
It has long been known that bacteria not only cause diseases, but are also widely used in biotechnology. In addition to long-established applications, e.g., for the bacterial production of proteins, appropriately configured bacteria have recently become increasingly important in the diagnosis or even treatment of diseases.
In the future, an approach is conceivable in which bacteria are directed to the right places in the body so that they can release active substances or produce proteins in a targeted manner after activation.
In order to be able to use bacteria in this way, targeted activation using red light, which can penetrate living tissue, is a good option. However, this requires an understanding of the fundamentals of molecular responses to external stimuli such as light.
With its study, the research team from the Photobiochemistry working group at the University of Bayreuth is contributing to the understanding and development of new approaches in the activation of bacteria by red light. The group is also providing fundamental principles in the sensory perception of bacteria.
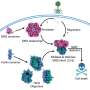
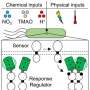
Bacteria must constantly adapt to changing external signals such as temperature, pH value or light. At the molecular level, these adaptations are often made by adding or splitting off phosphate groups. In many bacteria, a two-component system consisting of a light-sensitive enzyme and a regulator is often responsible for these processes.

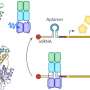
In some systems, the enzyme cleaves phosphate groups from the regulator under red light; in the dark, the enzyme adds phosphate groups to the regulator. The regulator then triggers further molecular processes in the bacteria, such as a change in gene activity, which can be used to produce almost any protein.
Researchers at the University of Bayreuth have modified a bacterial two-component system and thus shown that bacterial systems can be specifically reprogrammed in their physiological response to external stimuli.
For their model system, Stefanie Meier, a doctoral student in the Photobiochemistry working group, and Prof. Dr. Andreas Möglich, head of the working group, replaced the light-sensitive unit of the two-component system with another one. This made the two-component system ten times more sensitive to red light than the original variant.
The researchers found the reason for the higher sensitivity to light in the altered activity of the phosphate groups: In the modified two-component system, the phosphate groups were split off more quickly under red light compared to the original system. This means that the system is inactivated even at low red light intensities.
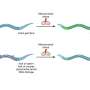
The researchers also changed the length of the connection (the "linker") between the light-sensitive unit and the rest of the enzyme. They found that the systems with modified linkers exhibited contrasting properties in light regulation and signal response at the genetic level to the original systems.
In the eyes of the researchers, the resulting variants with increased sensitivity and reprogrammed activity serve as novel tools for applications in synthetic biology and biotechnology.
"The results obtained from this model system have general relevance for countless systems of this type, which regulate important bacterial responses such as development, movement and infectivity. In addition, we are creating systems that can be used directly in biotechnology, which allow the production of any proteins to be activated by red light," says Möglich.
More information: Stefanie S. M. Meier et al, Leveraging the histidine kinase-phosphatase duality to sculpt two-component signaling, Nature Communications (2024). DOI: 10.1038/s41467-024-49251-8
Journal information: Nature Communications
Provided by Bayreuth University