This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers devise new way to find proteins for targeted treatment of disease
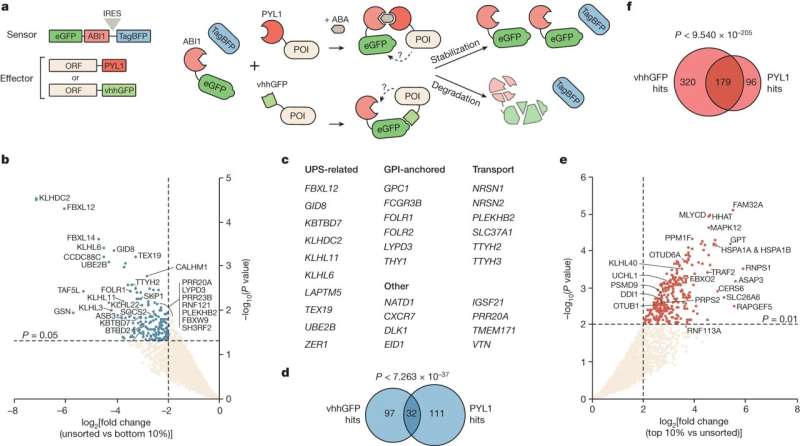
Researchers at the University of Toronto and Sinai Health have created a new platform to identify proteins that can be co-opted to control the stability of other proteins—a new but largely unrealized approach to the treatment of disease.
The researchers developed a method to interrogate the entire human proteome for 'effector' proteins, which can influence the stability of other proteins via induced proximity. The study marks the first time researchers have searched for effector proteins on this scale and has identified many new effectors that could be used therapeutically.
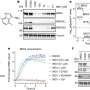
"We found more than 600 new effector proteins in 14,000 genes," said Juline Poirson, first author on the study and visiting scientist at U of T's Donnelly Centre for Cellular and Biomolecular Research. "Over 200 of the new effectors can efficiently degrade their target proteins, while about 400 effectors were capable of stabilizing, and thereby increasing the abundance of, an artificial target protein."
The study, which involved researchers at Sinai Health's Lunenfeld-Tanenbaum Research Institute, was published in the journal Nature.
"Targeting proteins through induced proximity is a new and promising area of biomedical research," said Mikko Taipale, principal investigator on the study and an associate professor of molecular genetics at the Donnelly Centre and the Temerty Faculty of Medicine.
"Not only did we find new effectors worth further investigation for drug discovery, we developed a synthetic platform that can be used to conduct unbiased, proteome-wide, induced-proximity screens to continue expanding the library of effector proteins."
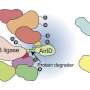
The effectors currently in use for targeted protein degradation and stabilization are E3 ubiquitin-ligases (E3s) and deubiquitinases (DUBs), respectively. E3 is an enzyme that transfers the ubiquitin molecule to the target protein, which essentially flags the protein for a proteosome to digest it. On the other hand, a DUB enzyme removes the ubiquitin tag from a protein, thereby preventing the protein from being recognized and degraded by a proteosome.
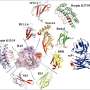
The results of the study demonstrate that E3s are quite varied in the degree to which they can degrade target proteins they are brought into contact with. The research team even discovered four of what they call 'angry E3s,' which consistently degrade targets regardless of other factors, such as the location of the target within the cell.
A particularly surprising finding was that some of the strongest effectors for targeted protein degradation were E2 conjugating enzymes instead of E3s. These differ from E3s in that they are involved at an earlier step of protein degradation and do not directly engage the target protein.
Because E2s were not considered to be easily druggable, they had not been harnessed for targeted protein degradation until recently. They represent, however, the untapped potential of stronger effectors than ones currently in use.
The study shows that exploring the whole proteome for induced proximity offers enormous opportunities for therapeutic interventions. KLHL40, one of the identified effectors, could potentially be hijacked for targeted protein stabilization to treat skeletal muscle disorders. The research team also found that targeted protein degradation with FBXL12 and FBXL15 effectors could be particularly useful in treating chronic myeloid leukemia.
Targeted protein degradation and stabilization are innovative methods of drug discovery that have thus far been plagued with the "protein pair problem," where the best effector for a target protein cannot be predicted accurately. Matching a target protein with the right effector is essential to successfully, and safely, facilitate degradation and stabilization processes in tissues.
"The synthetic screening platform developed by our team solves the protein matching issue through rapid, large-scale testing of effector and target protein interactions," said Poirson. "We're confident that an unbiased induced-proximity approach can be used to find effectors for almost any target."
More information: Juline Poirson et al, Proteome-scale discovery of protein degradation and stabilization effectors, Nature (2024). DOI: 10.1038/s41586-024-07224-3
Journal information: Nature
Provided by University of Toronto