This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
New class of 'intramolecular bivalent glue' could transform cancer drug discovery
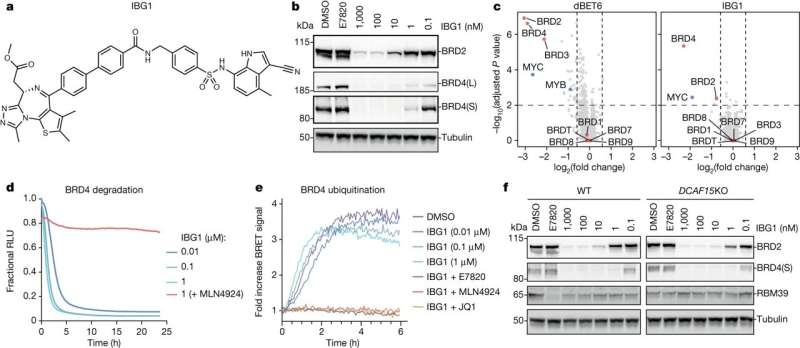
A breakthrough class of molecular glue identified at the University of Dundee could pave the way for a new generation of drugs to target cancers and neurodegenerative diseases.
A research team at the University's Centre for Targeted Protein Degradation (CeTPD) led by Professor Alessio Ciulli, in collaboration with the research group of Dr. Georg Winter at the Research Center for Molecular Medicine (CEMM) of the Austrian Academy of Sciences in Vienna, have defined a new class of so-called "intramolecular bivalent glue," which bind proteins—crucial to the cells that allow our bodies to function correctly—that would otherwise stay apart.
This research has been published in the journal Nature.
"These findings have major implications for the entire pharmaceutical industry engaged in targeted protein degraders," said Professor Alessio Ciulli, Director of Dundee's CeTPD.
"This is particularly true for the development of drugs that target cancer, neurodegenerative diseases, and many more illnesses driven by proteins that have always been considered undruggable."
"Proteins are essential for our cells to function properly, but when these do not work correctly, the body is vulnerable to disease."
"The glue that we have been able to define is special because it first attaches itself to one protein in two places—not just one—and then recruits the second protein, effectively sandwiching the two proteins together."
"We have only been able to identify this using our Targeted Protein Degradation technology and have identified a vulnerability that can be exploited by the design of new drugs that could potentially transform treatment for cancer patients and those with other untreatable diseases."
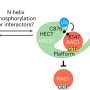
Targeted protein degradation (TPD) is an emerging field of drug development for treating diseases that involves redirecting protein recycling systems in our cells to destroy disease-causing proteins. Most TPD strategies use small molecules—so-called degraders—to recruit these target proteins to a class of enzymes called ubiquitin E3 ligases.
The E3 tags the target protein with ubiquitin labels, which ultimately leads to the destruction of the disease-causing protein via the cellular waste bin: the proteasome.
Working with collaborators at CEMM, the Goethe University of Frankfurt, and Eisai Co. Ltd, the Japanese pharmaceutical company, the Dundee team has been able to unveil a novel mechanism of molecular gluing, different from those previously known. This new mechanism binds to two sides of the target protein instead of just one, prompting a rearrangement of the whole protein and stabilizing its previously unknown interaction with the E3 ligase.
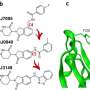
Furthermore, the team was able to visualize, for the first time, the precise mechanism by which their compounds work and bring together the target proteins to one of these E3 ligases. Because the molecules have two heads, which latch on to two different regions within the same target protein, these have been coined "intramolecular bivalent glues."
This world-leading work has also illuminated previously underappreciated features and properties of molecular glues, paving the way for scientists to develop a deeper understanding of glues that could allow for new classes to be discovered more rapidly.
"The impact of what we have revealed here cannot be underestimated," added Professor Ciulli. "This will cause a ripple effect throughout the pharmaceutical industry and has the potential to transform how we view drug development. I must also pay tribute to our collaborators, whose input has been crucial in achieving this seismic breakthrough."
More information: Alessio Ciulli, Targeted protein degradation via intramolecular bivalent genes, Nature (2024). DOI: 10.1038/s41586-024-07089-6. www.nature.com/articles/s41586-024-07089-6
Provided by University of Dundee