This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Uncovering how HIV assembles its lipid coat, which allows it to enter cells
New insights into how the human immunodeficiency virus (HIV) curates and assembles its lipid envelope have been gleaned by RIKEN biologists. These findings into HIV biology could help to inform the search for new treatments. The paper is published in the journal Nature Communications.
A well-chosen outfit can open doors in the human world. The same holds true for HIV, which wraps itself in a coat of specialized lipids that greatly affects its ability to gain an entry into—and subsequently exit—host cells.
Like most viruses, HIV has only a bare-bones set of essential genes and lacks virtually all of the metabolic functions present in cells.
"Since viruses cannot synthesize lipids, HIV 'steals' lipids from the plasma membrane of host cells," explains Toshihide Kobayashi of the RIKEN Pioneering Project on Integrated Lipidology.
However, the biochemical composition of this membrane differs notably from that of the viral envelope, which is heavily enriched for two subtypes of lipids: cholesterol and sphingomyelin.
A viral protein called Gag facilitates the gathering of these lipids as newly replicated viruses "bud" from the membranes of infected cells, but the underlying mechanism was unclear.
To fathom the depths of this mystery, Kobayashi teamed up with researchers at the University of Strasbourg in France, where he also maintains a lab.
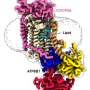
The expression of the HIV Gag protein is sufficient to drive plasma membrane budding in cultured human cells, and the team used a sophisticated multipronged imaging strategy to observe this process.
"Our lab has been developing and characterizing proteins that bind to specific lipids," says Kobayashi. The team labeled sphingomyelin- and cholesterol-binding proteins with fluorescent dyes. They then observed how these lipids behave during Gag-induced budding with cutting-edge microscopy methods that can resolve single molecules.
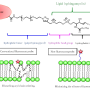
The plasma membrane consists of two layers: the inner and outer leaflets (named based on their position relative to the cellular interior).
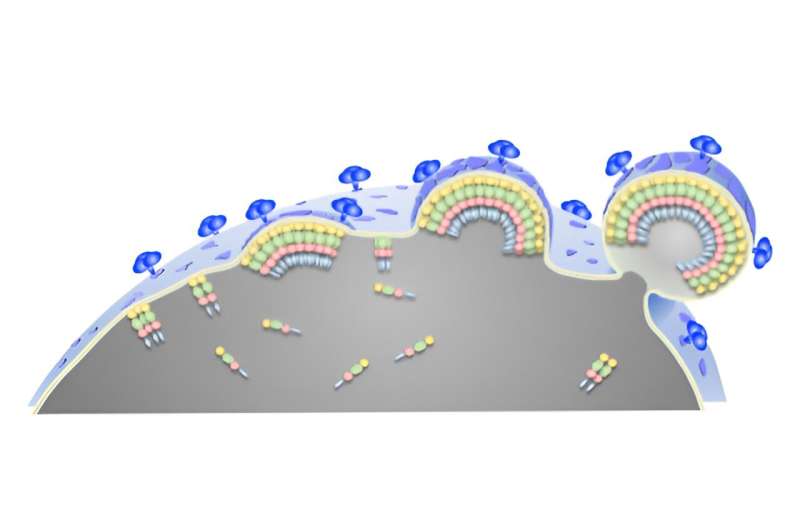
The researchers observed that when Gag attached to the inner leaflet, it overlapped with islands of cholesterol and sphingomyelin on the outer leaflet, physically isolating these lipids. As more Gag proteins bound the leaflet and interacted with each other, sphingomyelin accumulation continued to increase.
As a consequence, the nearby surface of the membrane began to curve, further concentrating these lipid domains as a prelude to the final stages of the budding process.
These findings represent an important step forward to understanding HIV biology, but key questions remain. "We've shown that the inner leaflet protein reorganizes outer leaflet lipids, but we do not know how," says Kobayashi. "Our priority is to clarify this mechanism."
More information: Nario Tomishige et al, HIV-1 Gag targeting to the plasma membrane reorganizes sphingomyelin-rich and cholesterol-rich lipid domains, Nature Communications (2023). DOI: 10.1038/s41467-023-42994-w
Journal information: Nature Communications
Provided by RIKEN