This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
From infamy to ingenuity—bacterial hijack mechanisms as advanced genetic tools
Researchers have uncovered the intricate molecular mechanism used by parasitic phytoplasma bacteria, known for inducing "zombie-like" effects in plants. This detailed revelation opens new horizons for groundbreaking applications in biotechnology and even in biomedicine.
The team led by Professor Saskia Hogenhout at the John Innes Centre, in partnership with The Sainsbury Laboratory, has employed X-ray crystallography to unveil the structure and functional mechanism of SAP05. This molecule plays a crucial role in bridging two distinct components inside plant cells.
The discovery sheds new light on a peculiar phenomenon in nature—mostly seen in "witches' brooms" in which plant stems and leaves proliferate due to Phytoplasma bacteria.
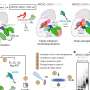
This insect-transmitted bacteria triggers diseases like Aster Yellows, significantly diminishing yields in leaf crops including oilseed rape, lettuce, carrots, grapevines, onions, and a variety of ornamental and vegetable crops worldwide.
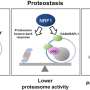
Previous research by the Hogenhout group revealed how the bacterial protein SAP05 is able to manipulate plants by hijacking molecular machinery called the proteasome. The proteasome breaks down and recycles proteins that are no longer required inside plant cells.
SAP05 hijacks this process, causing proteins that regulate growth and development to be dispensed into a molecular recycling center known as the 26S proteasome.
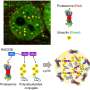
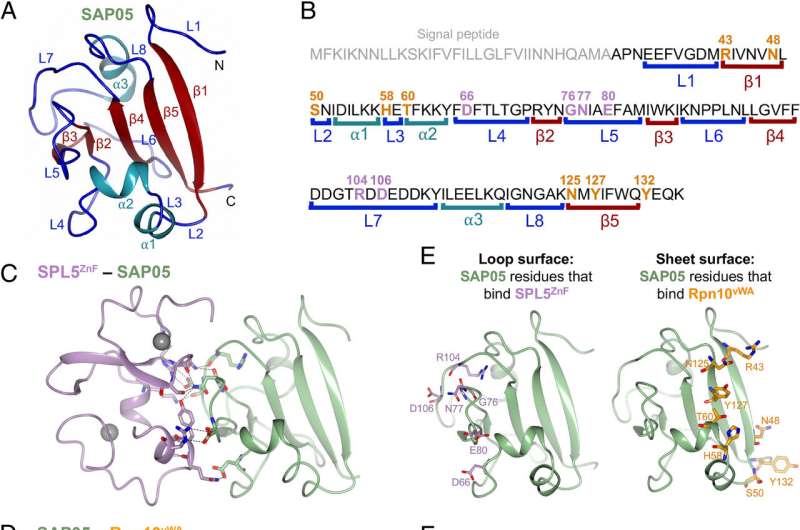
This latest research focuses on how this happens at the structural level. SAP05 effectively disrupts the molecular recycling pathway, serving as a scaffold that connects its two cellular targets: a transcription factor and the proteasome.
Fascinatingly, SAP05 binds in a manner that enables it to 'lift the dustbin lid," selectively disposing of developmental proteins, while strategically preserving functions vital for the survival of its plant host.
Professor Hogenhout, group leader at the John Innes Center and lead of the research team behind these findings, explains, "We now know the structure of this complex and how the protein binds to two cellular components to create a short circuit. While SAP05 allows itself to get involved in the plant, it does not disrupt other important processes. It is so amazing to see evolution crystallized in this way."
Usually, in plants, in fact, across all multicellular organisms, this recycling of proteins in the proteasome is dependent on a molecule called ubiquitin.
By short-circuiting this process, SAP05 provides a new way of carrying out the essential task of protein degradation, which serves its own parasitic purpose and is completely independent of ubiquitin.
The discovery presents some intriguing possibilities. The researchers were struck by the sophisticated ingenuity of SAP05, a master manipulator, and its promising applications in biotechnology.
First author of the paper Dr. Qun Liu says, "It was so exciting to see that this molecule SAP05 had two sides, one side binding to the transcription factor and the other binding to the 26S proteasome. It's very smart."
By understanding how this bacterial mechanism interacts with cells at a structural level, the researchers can now use this knowledge to engineer SAP05-like molecules which could be repurposed to remove unwanted proteins such as pathogen effectors or viruses, with an impact on therapeutics, research, and in agriculture.
The study is published in the journal Proceedings of the National Academy of Sciences.
More information: Qun Liu et al, Bimodular architecture of bacterial effector SAP05 that drives ubiquitin-independent targeted protein degradation, Proceedings of the National Academy of Sciences (2023). DOI: 10.1073/pnas.2310664120
Journal information: Proceedings of the National Academy of Sciences
Provided by John Innes Centre