This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
New model helps researchers get a handle on unpredictable cells
In the process of embryonic development, cells take on intricate forms and functions. Patterns emerge from groups of cells, giving the biological tissues form, function, and integrity. It's a complex process that isn't well understood.
A new study, led by Prof. Yimin Luo, presents new methods to get a clearer picture of the cellular dynamics at play, which could provide a better understanding of many developmental processes, as well as how cancer cells invade tissue. The results are published in PRX Life.
Cells are a form of active matter—that is, a system of individual agents that interact with one another and consume energy from their surroundings, but collectively they form a much larger structure. It's a category that includes a wide range of biological systems, such as flocks of birds, schools of fish, monolayers of cells, colonies of bacteria, and others, all exhibiting intriguing phenomena.
But there's a big difference in how different systems of active matter behave. The smaller things get, the more mysterious they become. Unlike flocks of birds or schools of fish, it's much harder to predict the patterns of cells. While the patterns of bird flocking can be delineated clearly, cell trajectories are easily blurred by the forces in their environments.
"When things are much smaller, they're subjected to more random fluctuations," said Luo, assistant professor of mechanical engineering and materials science. "When you try to identify cellular motions, you usually do it under the microscope. These interactions are extremely complex because cells, they deform and then they have biochemical interactions with other cells, and they move very slowly."
That's only one complication. Another is the sheer number of cells that Luo's lab is observing and imaging.
"Most people image about a couple hundred, while we do a couple of thousand," she said. "So then the second curse and blessing is that if you have lots of observations, you can somehow offset the uncertainty in your observation. Technically, the more you observe, then the more you can isolate the signal from the noise."
But calibrating your model with an extremely large data set is very difficult, Luo said.
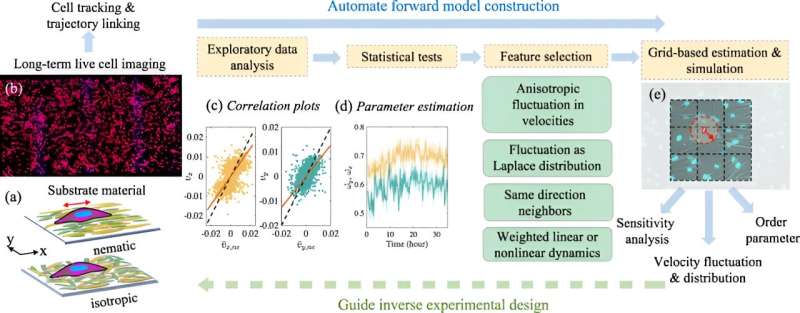
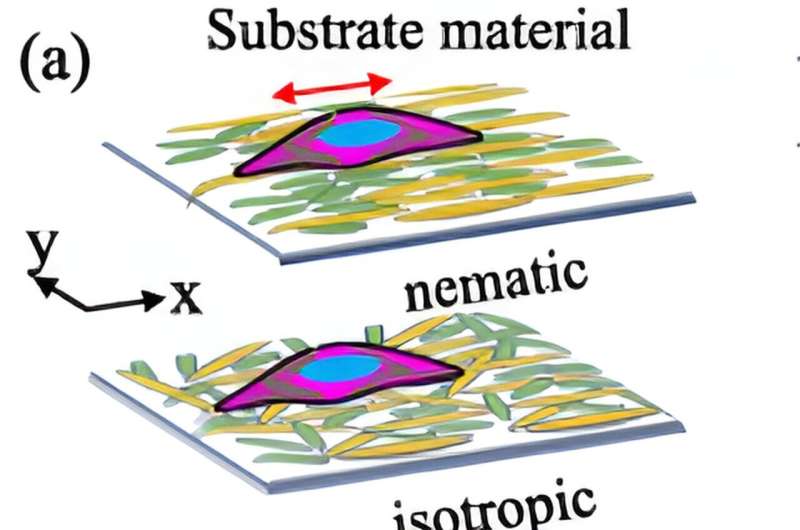
Cells align to form tissues mainly when they reach a very high density. To accurately model this, researchers need to take into account both the cell-to-cell interactions as well as the influence of their surrounding environments. To do so, Luo and her team developed a hybrid procedure that combines statistical learning approaches with physics-based models. The result is a more efficient method that avoids testing all feature combinations through simulations.
"We came up with this approach where we use a data-driven way to select features called the exploratory data analysis (EDA)," she said, adding that the process has been in use for decades. "With EDA, you work with the data to see what it tells you, and then you try to implement that into a model."
By computationally including key features of the cells' behavior, such as certain fluctuations and particular types of cell-to-cell interactions, the researchers can more accurately calibrate their physics-based models. Further, the computational tools they developed for automating model construction and calibration can be applied to other systems of active matter.
"It's a very general framework," she said. "It's an approach that's more flexible than physics-based models, but more rational and less labor-intensive than a purely data-driven approach."
More information: Mengyang Gu et al, Data-Driven Model Construction for Anisotropic Dynamics of Active Matter, PRX Life (2023). DOI: 10.1103/PRXLife.1.013009
Journal information: PRX Life
Provided by Yale University