This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
Team develops all-species coronavirus test
In an advance that will help scientists track coronavirus variants in wild and domesticated animals, researchers report they can now detect exposure to the SARS-CoV-2 virus in any animal species. Most coronavirus antibody tests require specialized chemical reagents to detect host antibody responses against the virus in each species tested, impeding research across species.
The virus that causes COVID-19 in humans also infects a variety of animals, said University of Illinois Urbana-Champaign pathobiology professor and virologist Ying Fang, who led the new research. So far, the virus has been detected in cats, dogs, rodents, deer, apes and a variety of farm and zoo animals. The virus also mutates in these hosts, potentially leading to new variants that can endanger their—and human—health.
"Highly sensitive and specific diagnostic reagents and assays are urgently needed for rapid detection and implementation of strategies for prevention and control of the infection in animals," the researchers wrote in the journal mSphere, where their findings are reported.
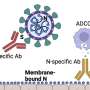
The new coronavirus test focuses on antibodies against a protein, called the N-protein, that is embedded in the virus's nucleocapsid—a structure made up of proteins and nucleic acids contained within a viral membrane. The N-protein makes a better target than the membrane-bound viral proteins that are usually used in tests for antibody responses, Fang said.
"The N-protein is more abundant and it is more conserved than the proteins used in most tests," she said. This means that the structure of the protein is more consistent across species, making it a good target for all-species antibody tests.
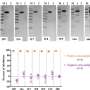
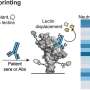
The team used an N-protein-based blocking ELISA protocol for their test. This method involves coating an ELISA plate with the N-protein, then adding a serum sample of whatever animal is being tested. If the animal has been infected with the coronavirus, its serum will contain anti-N-protein antibodies, which will bind to the N-protein-coated plate.
The scientists then wash the plate and add a secondary biotin-tagged monoclonal antibody that targets the N-protein. If the animal is positive for coronavirus infection, its antibodies will block the secondary antibodies from binding to the N-protein. If the animal has not been infected, the monoclonal antibodies will attach to the coated plate and generate a color signal when specific chemicals are added to the plate.
The researchers validated their test using samples from various animals with known SARS-CoV-2 infection status, finding the tests had more than 97% sensitivity and 98% specificity. Further tests in domestic cats showed that the assay was able to detect infection within seven days of exposure to the virus.
The development of accurate cross-species coronavirus tests provides a useful tool for SARS-CoV-2 field surveillance in animal populations, helping scientists identify potential new animal reservoirs to prevent future disease outbreaks, Fang said.
More information: Ying Fang et al, Development of monoclonal antibody-based blocking ELISA for detecting SARS-CoV-2 exposure in animals, mSphere (2023). DOI: 10.1128/msphere.00067-23 , journals.asm.org/doi/10.1128/msphere.00067-23
Provided by University of Illinois at Urbana-Champaign