This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
A glimpse into the hexasome: 40 years on
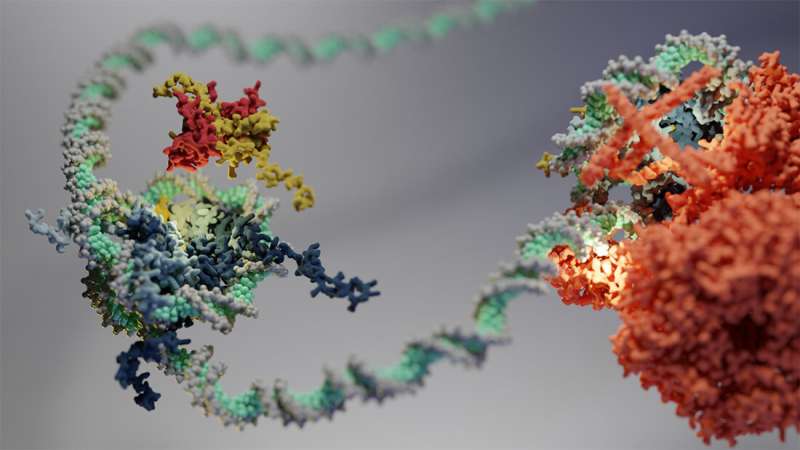
In 1983, scientists discovered hexasomes—a unique molecular structure that helps cells package their DNA. Now, a study conducted by the Eustermann group at EMBL Heidelberg has shed light on how DNA packaging into hexasomes can affect the function of enzymes involved in gene regulation. The research is published in the journal Science.
DNA is a very long, thin thread containing our genetic instructions. Being much longer than the tiny space inside our cells, it needs a clever packaging system. That's where nucleosomes come into play—tiny spool-like structures that help compact our genetic information.
Multiple nucleosomes are then linked together in a "beads-on-a-string" arrangement, collectively making up chromatin—a compact fiber comprising genetic material (DNA) and the proteins that help package and organize it (e.g. histones).
Nucleosomes also play a crucial role in regulating gene activity, determining which genes are switched on or off. The Eustermann group at EMBL Heidelberg aims to understand how these structures work in the context of DNA packaging.
"We are interested in understanding the molecular principles by which DNA is packaged in the nucleus of a eukaryotic cell," said Sebastian Eustermann, Group Leader at EMBL Heidelberg. "Our focus lies specifically on visualizing the molecular processes of how this packaging then regulates the expression and maintenance of the genome."
Nucleosomes and hexasomes under the cryo-EM lens
Nucleosomes can exist in different structural versions, known as non-canonical forms, which may play a fundamental, yet so far largely unexplored, role in genome regulation. The hexasome is one such version. A standard nucleosome wraps DNA around eight histone proteins, while a hexasome uses only six. Such small structural differences can play a major role in the molecular realm of the cell.
In recent years, thanks to breakthroughs in cryo-electron microscopy (cryo-EM), researchers were finally able to solve the structure of canonical nucleosomes along with their interacting proteins, providing unprecedented insights into how they collectively regulate the genome. However, until now, the molecular details of hexasomes and their interactors remained unclear—a gap Eustermann's team has been trying to fill.
"Forty years ago, hexasomes were discovered by my Ph.D. mentor Daniela Rhodes. Now, my team could finally obtain first insights into how hexasomes impact the function of enzymes called chromatin remodelers," said Eustermann.
Chromatin remodeling enzymes are crucial for maintaining the dynamic nature of chromatin packaging.Their activity can change chromatin structure and accessibility, leading to alterations in gene expression patterns.
"These enzymes work like molecular machines. Fuelled by ATP, an energy-rich cellular compound, they can spool DNA around histones," explained Anna Jungblut, co-author of the study and Ph.D. student in the Eustermann group. "Remodelers play a vital role in regulating the genome and have significant effects on cell function. For example, when gene regulation is disrupted, it frequently contributes to diseases like cancer."
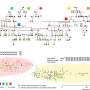
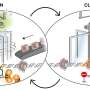
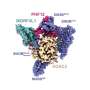
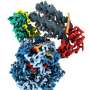
To investigate this process, the team made extensive use of cryo-EM, which allowed them to look at the atomic structure of a hexasome bound to a chromatin remodeler. They found that remodelers recognize and modify the structure of hexasomes, indicating that these non-canonical nucleosomes not only affect the way DNA is packaged, but also impact how enzymes interpret the regulatory information within chromatin.
"Our study uncovers how certain features of hexasomes can activate and regulate chromatin remodeling enzymes. These intriguing insights may inspire studies on other molecular machines in the nucleus of a cell," said Min Zhang, first author of the study and EIPOD postdoc in the Eustermann group. "Hexasomes are likely to have a similar impact on human remodeling enzymes closely linked to cancer. Follow-up studies on the regulation of hexasomes on the genome would provide new perspectives in gene regulation."
The discovery is further bolstered by findings from Yifan Cheng's and Geeta Narlikar's research groups at UCSF, who have shown structurally that chromatin remodeling enzymes become activated on hexasomes, even though hexasomes lack features that were previously thought to be essential for the enzymes to function. In an earlier study, the Narlikar group had demonstrated biochemically the link between hexasomes and chromatin remodeling.
"When Geeta Narlikar and I met at a conference last year, we were astonished that we independently made the discovery of how remodelers achieve their function on hexasomes," said Eustermann. "Again an example of how we can benefit from open exchanges and collaborative discussions in science."
More information: Min Zhang et al, Hexasome-INO80 complex reveals structural basis of noncanonical nucleosome remodeling, Science (2023). DOI: 10.1126/science.adf6287
Journal information: Science
Provided by European Molecular Biology Laboratory