This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
New insight into how xeroderma pigmentosum causative gene products ensure the accuracy of DNA repair
Our genomic DNA is continuously damaged by endogenous factors such as reactive oxygen species, and also by environmental factors such as ultraviolet light, radiation, and chemicals. Failure to repair damaged DNA may induce mutations and cell death, eventually leading to the onset of cancer and other diseases. To prevent this, our cells are equipped with various defense systems aimed at finding and repairing damaged DNA.
Nucleotide excision repair (NER) is an important mechanism for the repair of diverse DNA lesions caused by ultraviolet light and chemical carcinogens. Many patients diagnosed with xeroderma pigmentosum (XP) have mutations in one of the genes encoding proteins involved in the NER mechanism. It is known that their high susceptibility to ultraviolet light-induced skin cancer in particular is due to the deficiency in NER function. This also tells us that NER serves as a defense against cancer.
The first step of NER involves the XPC protein, one of the several XP causative gene products, which operates as a "sensor" to locate abnormalities in the DNA structure. XPC has a potential to deal with a wide variety of DNA lesions, while it also binds to DNA sites without any damage, such as base mismatches. Since unnecessary repair reactions at such improper sites may increase a risk of adverse mutations, it is essential to verify whether there is any damage that must be repaired. Although it has previously been shown that the basal transcription factor TFIIH and XPA protein play a role in such a damage verification step, the detailed mechanism has remained unclear up to now.
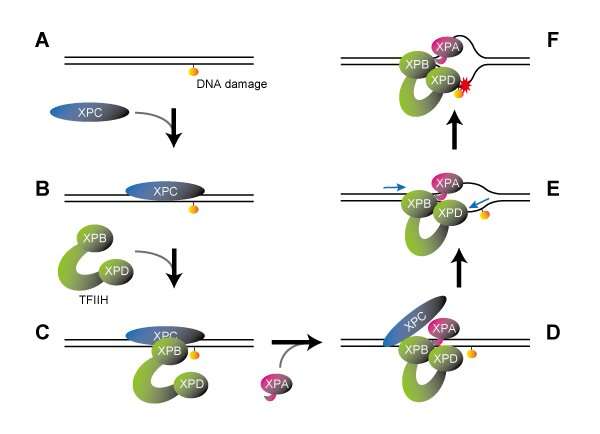
Following the known reaction steps in the NER process, researchers of a study published in Nature prepared a series of complexes containing damaged DNA sequentially bound by XPC, TFIIH and XPA, which were then subjected to cryogenic electron microscopy to analyze their molecular structures in detail. TFIIH is a large protein complex composed of seven subunits, including XPB and XPD proteins, which plays crucial roles in both NER and transcription (gene expression). It usually adopts a U-shaped "horseshoe" structure, with the XPB and XPD proteins positioned at the tips of its two open arms.
This horseshoe structure was retained in the DNA-XPC-TFIIH complex, with XPB bound to XPC at the damage site, while XPD did not interact with either DNA or XPC. When XPA was subsequently involved, however, TFIIH underwent a major conformational change. While the two open arms of the horseshoe came together to form a ring-shaped structure, XPB moved away from the damaged site such that XPD was allowed to bind DNA in between.
How does this molecular arrangement then verify the presence of DNA damage? It is known that XPB functions as a "DNA translocase," which binds to the DNA duplex and move along one of the DNA strands. Based on the newly elucidated structure for the complex, it was assumed that XPB should move along the DNA away from the damage site. However, since it is constrained by other proteins, XPB itself cannot actually move. Instead, the DNA duplex moves in the opposite direction. In other words, XPB pushes DNA into the complex.
Because the structure revealed that part of XPA is inserted between the two DNA strands behind XPB, it was expected to unwind the DNA duplex into single strands like a fastener being unzipped. Moreover, it was also known that XPD binds to single-stranded DNA and moves along it in a defined direction (5'→3').
Of the two DNA strands unraveled by XPB and XPA, XPD binds to the damaged one. However, as is the case with XPB, the XPD itself cannot move, so the damaged strand is likewise pulled towards the complex. When bound to a single-stranded DNA, a narrow pinhole is formed in XPD, through which the DNA strand passes. Therefore, if there is a bulky lesion on the DNA strand, that part will be trapped at the entrance to XPD such that the strand will stop its moving (like a knotted thread being unable to pass through the eye of a needle.
The present research thus successfully elucidated the mechanism whereby the presence of a DNA lesion is verified: The decision to proceed with the repair reaction is made by the blockage of DNA strand movement through XPD. In fact, previous research has indicated that non-bulky DNA lesions (base detachment, etc.) that could be predicted to pass through the XPD hole are not a target for the NER repair process.
The current research has clarified which parts of proteins such as XPA, XPB, and XPD are involved in the NER process to repair damaged DNA together with their mechanism of action. As our understanding deepens about the negative impact on repair response arising from structural changes induced in pathogenic variants of these proteins in XP patients, it may also be possible to support the future development of drugs and other treatments.
More information: Jinseok Kim et al, Lesion recognition by XPC, TFIIH and XPA in DNA excision repair, Nature (2023). DOI: 10.1038/s41586-023-05959-z
Journal information: Nature
Provided by Kobe University