This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers shed new light on the motor of DNA replication
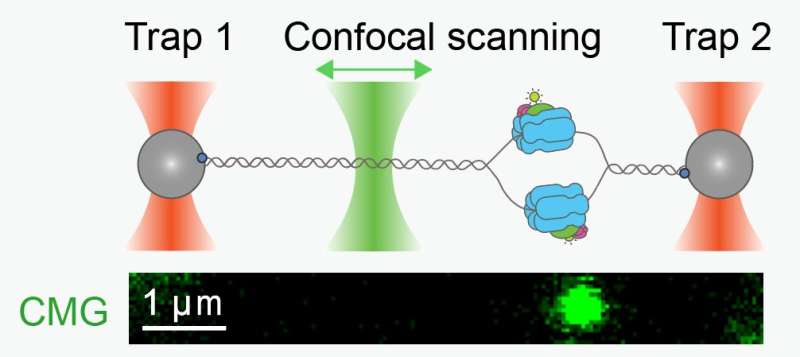
DNA replication is the process whereby cells make an exact copy of their DNA before cell division. A key part of the intricate DNA replication machinery is a molecular motor called CMG, which has the vital task of separating the two strands of the DNA double helix so that they can be copied.
An interdisciplinary team of researchers from TU Delft has now developed a new methodology to assemble and image the motion of CMG with unprecedented resolution. Their findings, published on April 14, 2023 in the open-access journal Nature Communications, will pave the way for further discoveries in the study of DNA replication.
DNA replication is fundamental to life, as the faithful transmission of genetic information between generations of cells is crucial for the survival and health of all living organisms. It is carried out by an intricate cellular machinery, known as the replisome, that is made up of protein building blocks. A key component of this machinery is CMG, the molecular motor that powers the replisome during DNA replication.
CMG has the important task of separating the two strands of the DNA double helix, so that the message that they encode can be read and copied. "Understanding how CMG moves along DNA is crucial for our understanding of DNA replication," says Daniel Ramírez Montero, researcher and first author of the publication. Studying DNA replication is very important because errors in this process can lead to genetic disorders or cancer.
Motion on film
In living cells, CMG is assembled and activated via an intricate cascade of biochemical reactions that involves 36 different proteins. A group of TU Delft researchers led by Spinoza laureate Professor Nynke Dekker, in collaboration with Francis Crick Institute group leader Dr. John Diffley, has developed a way to carry out this tightly controlled process outside of the cell and measure the motion of individual CMG molecular motors.
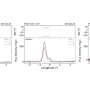
The researchers extracted all 36 proteins from cells to build up CMG on DNA. By attaching fluorescent labels to some of the proteins, they could directly observe the movement of the CMG molecular motor under a fluorescent microscope.
"With this new approach, we were able to visualize the motion of individual CMGs made from scratch. We used optical tweezers to hold the CMG-containing DNA still for easier visualization, and then took movies of CMG as it moved along the DNA. This way, we could measure its motion at the single-molecule level for the first time," explains Ramírez Montero.
New insights
Using their bottom-up approach, which combines cutting-edge biochemistry and biophysics, the group of researchers was able to directly visualize the motion of individual CMG motors assembled from scratch for the first time, and measure this motion with unprecedented resolution. In addition, they made the unexpected discovery that CMG can move randomly along the DNA when a key molecule called ATP is absent; furthermore, they show that subsequent re-binding of ATP allows CMG to tightly hold onto the DNA, thereby halting its random motion. This halting is important, as it likely facilitates the activation of CMG, a critical process in the initiation of DNA replication.
This work will pave the way for further studies that may uncover unknown details of key processes in DNA replication. These discoveries can in turn get us closer to understanding how cells manage to faithfully transmit their genetic information with every cell division, as well as to better understand the errors in the process that can contribute to the development of genetic disorders or cancer. States Ramírez Montero, "Biological systems can look very complicated and chaotic at first sight, but by observing them at this resolution, we can understand the simple and elegant physics behind them."
More information: Daniel Ramírez Montero et al, Nucleotide binding halts diffusion of the eukaryotic replicative helicase during activation, Nature Communications (2023). DOI: 10.1038/s41467-023-37093-9
Journal information: Nature Communications
Provided by Delft University of Technology