This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
New discovery dramatically reduces time it takes to build molecules

With a big assist from artificial intelligence and a heavy dose of human touch, Tim Cernak's lab at the University of Michigan has made a discovery that dramatically speeds up the time-consuming chemical process of building molecules that will be tomorrow's medicines, agrichemicals or materials.
The discovery, published in the Feb. 3 issue of Science, is the culmination of years of chemical synthesis and data science research by the Cernak Lab in the College of Pharmacy and Department of Chemistry.
The goal of the research was to identify key reactions in the synthesis of a molecule, ultimately reducing the process to as few steps as possible. In the end, Cernak and his team achieved the synthesis of a complex alkaloid found in nature in just three steps. Previous syntheses had taken between seven and 26 steps.
"Making a chemical structure that has atoms in just the right place to give you efficacious and nontoxic medicines, for instance, is tricky," said Cernak, assistant professor of medicinal chemistry and chemistry. "It requires a chemical synthesis strategy grounded in the chemical building blocks you can actually buy and then stitch together using chemical reactions."
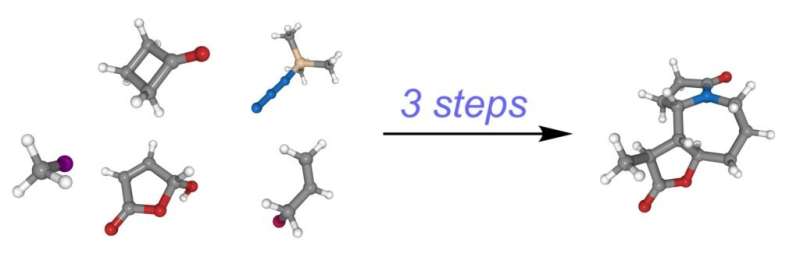
The accomplishment has powerful implications for speeding up the development of medicines.
Cernak compared the construction of these complex molecules to playing chess. You need to orchestrate a series of moves to get to the end of the game. While there's a near infinite number of possible moves, there's a logic that can be followed.
"We developed a logic here, based in graph theory, to get to the end as quickly as possible," he said.
Cernak and colleagues used SYNTHIA Retrosynthesis Software, which provides scientists with a database of pathways, or steps, and formulas for millions of molecular structures. This gave the team an enormous amount of computational synthesis data to play with.

Using an algorithm they developed to curate the data, the researchers identified the steps along the pathway that were high impact, or key steps, and the steps that were making progress toward completing the synthesis but ultimately inefficient for the whole process.
"We hope this research can lead to better medicines," Cernak said. "So far, we have been limited in the molecular structures we can quickly access with chemical synthesis."
Co-authors include Yingfu Lin, senior research fellow in pharmacy; Rui (Sam) Zhang, doctoral student in chemistry; and Di Wang, doctoral student in pharmacy.
More information: Yingfu Lin et al, Computer-aided key step generation in alkaloid total synthesis, Science (2023). DOI: 10.1126/science.ade8459
Journal information: Science
Provided by University of Michigan