This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Raman spectroscopy method for rapid identification of beer spoilage bacteria

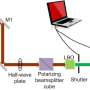
In a study published in Analytical Methods, a research group led by Li Bei from the Changchun Institute of Optics, Fine Mechanics and Physics (CIOMP) of the Chinese Academy of Sciences (CAS) proposed the rapid detection of beer spoilage bacteria based on label-free surface-enhanced Raman spectroscopy (SERS) technology.
Lactic acid bacteria are common spoilage bacteria in beer and need to be monitored and controlled at all stages of beer production. Traditional spoilage bacteria detection methods are time-consuming and cannot meet the demand for real-time, in-situ, rapid detection during the production process.
Raman spectroscopy has been widely used for microbial detection due to its fast, non-destructive and accurate properties, but conventional Raman spectroscopy has the disadvantage of poor signal-to-noise ratio, which affects the accuracy of microbial identification.
Compared with conventional Raman spectroscopy, the SERS technique has a stronger and more sensitive signal and is well suited to the detection of beer spoilage bacteria. Furthermore, the label-free SERS technique is ideal for commercialization due to its low cost and good results.
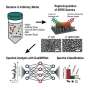
In this study, the researchers improved the existing process for the preparation of label-free SERS silver nanoparticles (AgNPs). The effect of the AgNPs@KCl agglomeration effect on SERS enhancement was investigated. Eight species of beer spoilage bacteria produced during the beer brewing process were identified by SERS.
The researchers further investigated the effect of the method on the final identification rate by combining the t-distributed stochastic neighbor embedding (t-SNE) dimensionality reduction analysis algorithm, Support Vector Machine (SVM), k-NearestNeighbor (KNN) and Linear Discriminant Analysis (LDA) machine learning algorithms. All three machine learning algorithms achieved an accuracy of around 90% and performed well in identifying beer spoilage bacteria.
In the stability analysis and mixing tests, two known spoilage bacteria were mixed with pure beer and incubated at constant temperature for a period of time to identify the bacteria in the beer. The two spoilage bacteria were successfully detected in the samples and had good spectral stability.
According to the final validation study, the technique can indeed identify the target spoilage bacteria from the simulated samples, which is of great significance to the rapid identification of spoilage bacteria in the beer brewing process.
More information: Lindong Shang et al, Rapid detection of beer spoilage bacteria based on label-free SERS technology, Analytical Methods (2022). DOI: 10.1039/D2AY01221A
Journal information: Analytical Methods
Provided by Chinese Academy of Sciences