This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Nanopore-based sensing device explores neurodegenerative diseases

Tau and tubulin proteins are two of the leading causes of many neurodegenerative diseases, such as Alzheimer's and Parkinson's. Most of neurodegenerative disease progression is related to the aggregation of these proteins in the brain.
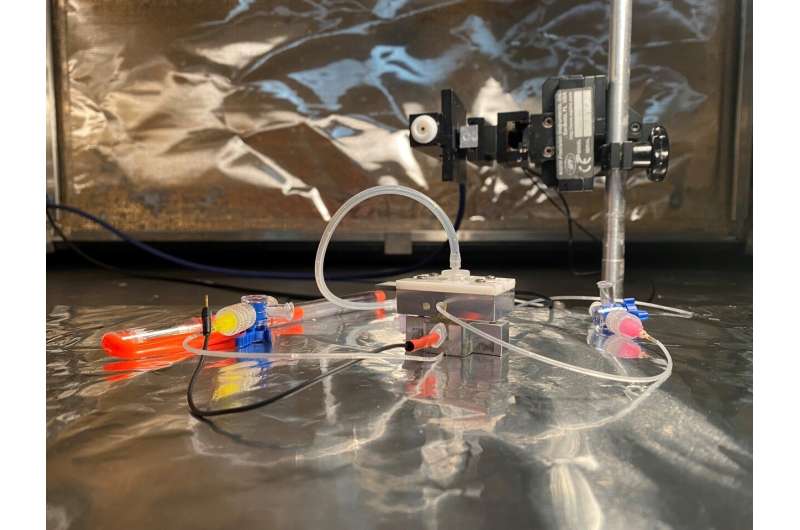
Inspired by one of her doctoral students who wanted to explore tau and tubulin proteins, Jiali Li, a physics professor at the University of Arkansas, and her group created a special silicon nitride nanopore-based sensing device.
In the Journal of Applied Physics, Mitu Acharjee and colleagues present the device, which is designed to provide volume information about tau and tubulin protein molecules and their aggregation states at the single-molecule level within their native environment.
To create the sensor, the team explored how the proteins change the current and voltage flowing through a nanopore system.
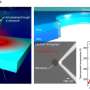
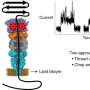
"Ohm's Law is the basic physics that enables the nanopore device to sense protein molecules," said Li. "A tiny hole—from 6 to 30 nanometers—is made in a thin silicon nitride membrane and supported by a silicon substrate. When that is placed into a solution with salt ions, applying an electric voltage drives the ions' flow through the hole, or nanopore. This, in turn, generates an open pore ionic current."
When a charged protein molecule—often thousands of times larger than the ions—is near the nanopore, it also gets driven into the nanopore and blocks the flow of some ions. This causes the open pore current to drop.
"The amount of current drop produced by a protein molecule is proportional to the protein's volume or size and shape," said Li. "This implies that if protein A binds to protein B, they will cause a current drop proportional to the volume of A+B, and an aggregated protein A will cause approximately multiple amounts of current drop."
This allows Li and her group to look at the protein binding and aggregation within a nanopore device. The amount of time a protein stays in a nanopore is inversely proportional to its charge, which also provides useful information about a protein molecule.
"Our study shows that a silicon nitride nanopore device can measure volume information of tau and tubulin protein molecules and their aggregation under different biological conditions, and this gives us a better understanding of the protein aggregation process, as well as developing drugs or other therapeutic methods to treat neurodegenerative diseases," said Li.
Using their solid-state nanopore device, along with other nanotechnology tools, "we plan to study the mechanism of protein aggregation under different biological conditions systematically, such as temperature, pH, and salt concentration," she said.
The article "Tau and tubulin protein aggregation characterization by solid-state nanopore method and atomic force microscopy" is authored by Mitu C. Acharjee, Haopeng Li, Ryan Rollings, Bo Ma, Steve Tung, and Jiali Li. It will appear in the Journal of Applied Physics on Jan. 10, 2023.
More information: Tau and tubulin protein aggregation characterization by solid-state nanopore method and atomic force microscopy, Journal of Applied Physics (2023). DOI: 10.1063/5.0123688
Journal information: Journal of Applied Physics
Provided by American Institute of Physics