January 27, 2023 feature
This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Fish-on-Chips: An optofluidic platform to investigate the neural and chemosensory axes of zebrafish
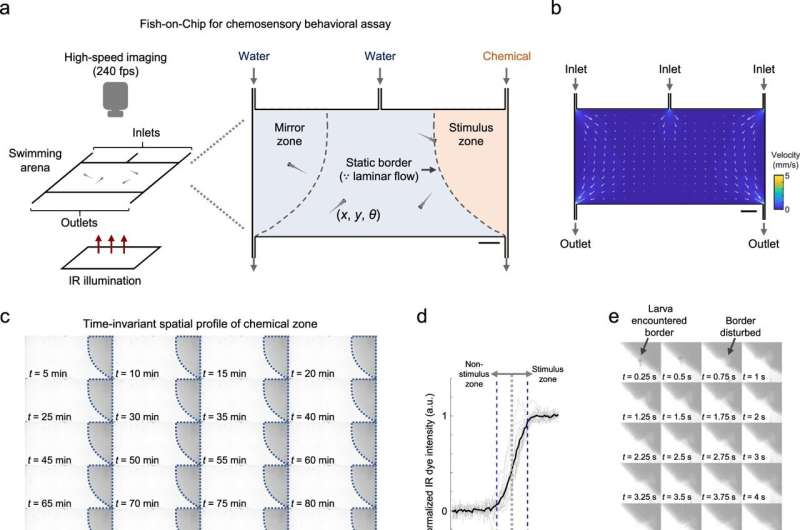
Neuroscientists study chemosensory processing by establishing chemical cues and the corresponding behavioral responses to record large-scale neuronal activity. In a new report now published in Nature Communications, Samuel Sy and a team of scientists in neurology, health sciences, biomedical engineering and mathematics in China and France presented a new method based on a set of optofluidic tools. This technology established chemical delivery to simultaneously image the behavioral outputs and whole-brain neural activities at cellular resolution in larval zebrafish.
The team included a fluidics-based swimming arena within the experimental setup and an integrated microfluidics-light sheet fluorescence microscopy system. The technical methods incorporated laminar fluid flow to achieve chemical cue representation. The team named the new technique "Fish-on-Chips," which is now ready to empower investigations of neural coding in the chemical senses.
Systems-level investigations of the brain
Neuroscientists explore the brain to understand behavioral goals of animals and comprehend the underlying algorithmic and neuronal mechanisms. Researchers have thus far developed a variety of sophisticated tools to mimic the natural habitats and thereby reconstruct the sensory environments of an animal for behavioral analysis at cellular resolution. For example, larval zebrafish are attractive vertebrates due to their capacity to rapidly acquire innate chemosensory, auditory and visually guided behaviors.
Evolutionarily, chemosensation is the oldest existing sensory system, and researchers have made considerable efforts to understand the behaviors underlying neural basis in a variety of popular model organisms in systems neuroscience.
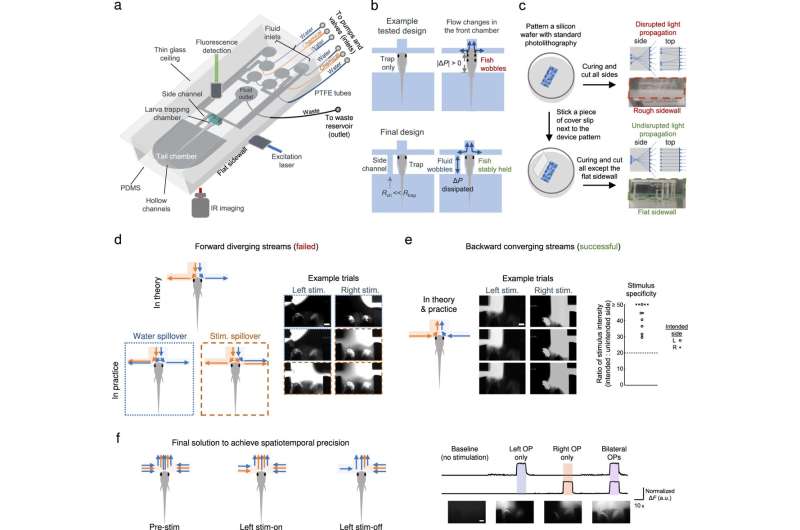
While fluidics and microfluidics provide precise fluid regulation to study chemosensation, a custom microfluidics-integrated light-sheet fluorescence microscope can establish whole-brain neuronal activity imaging at cellular resolution. In this report, Sy and colleagues therefore used a fluidics and microfluidics-based method alongside a microfluidics-integrated light sheet fluorescence microscope to study chemosensation for chemical cue delivery in small animals.
Chemosensory behavioral assays
To understand chemosensory-guided behavior under biomimicry, the team established a behavioral assay for larval zebrafish with a precisely defined arena in which the concentration profile of a chemical remained constant. The team maintained equal flow rates for all three inlets to establish clear separation of fluidic stream zones in the presence of actively swimming larvae.
Additionally, the researchers uncovered the circuit principles of chemosensory processing to deliver chemical stimuli while simultaneously recording neuronal activity. They accomplished this by stabilizing the larval subjects to establish compatibility with neuronal imaging methods while precisely regulating the delivery of a chemical stimuli. They also designed a light sheet fluorescence microscope-compatible microfluidic chip to stabilize the larvae for high-quality neuronal activity imaging and thereby ensured microfluidic chip compatibility.
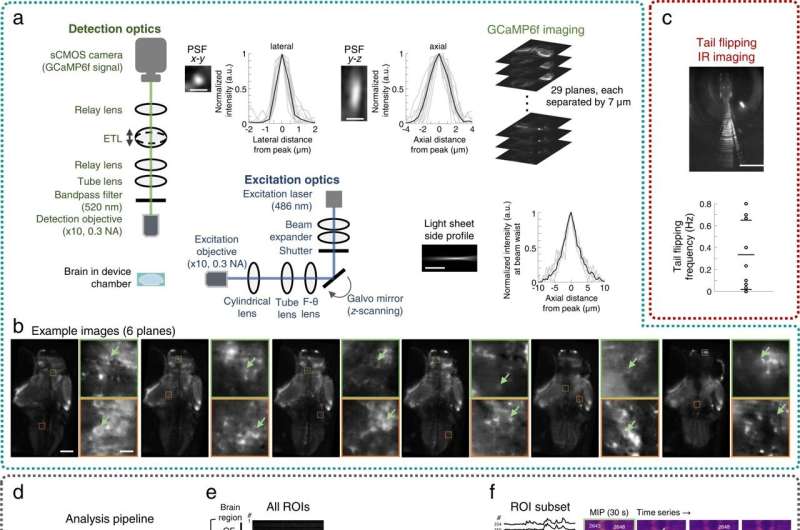
Imaging neuronal activities and behavior
The research team next sought to stably integrate the microfluidic components with a custom-built light sheet fluorescence microscope for additional experiments. Using the platform, they examined nine behaviorally active larvae with spontaneous tail flipping frequency for behavioral recording and chemosensory activities, as well as associated neuronal behavior at cellular resolution within the Fish-on-Chips navigation arena. The scientists also explored chemosensory behavioral algorithms underlying the species during cadaveric avoidance, where the model organisms relied on binasal inputs.
-
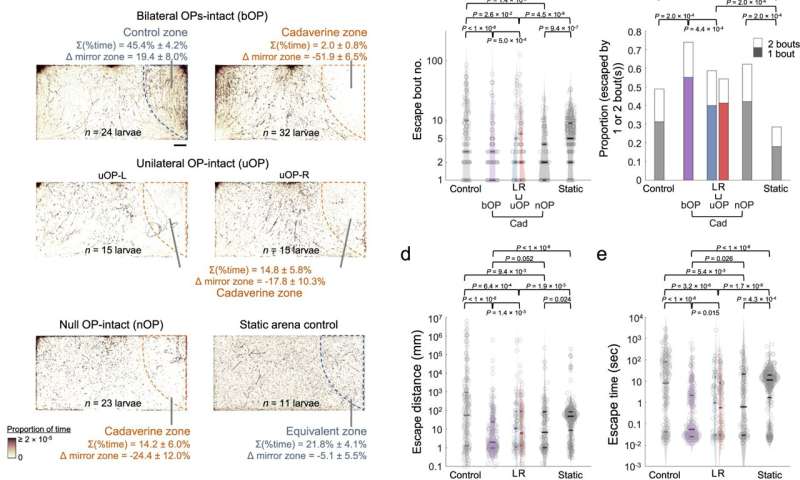
Larval zebrafish cadaverine avoidance revealed by the chemosensory behavioral assay. a) Upper panels: footprints of bilateral OP-intact (bOP) larvae in control and avoidance assays. Middle panels: footprints of unilateral OP-intact (uOP) larvae in avoidance assays. Lower panels: footprints of null OP-intact (nOP) larvae in avoidance assays and static arena control groups. Note that the static arena control group assays were carried out without flow. For each group, the rightmost (stimulus or water) zones are outlined by dashed lines. The percentages of time spent in the rightmost zone and their differences from that in the mirror water zone (Δ mirror zone) are shown (with SEMs across assays). Scale bar: 0.5 cm. b) Bout number, c proportion of entry-to-exit events with only 1 or 2 bouts (P value: one-sided Chi-squared test with Tukey's post-hoc test, comparing 2-bout event proportions), d distance traveled, and e time taken to escape the rightmost zone in control assays (Control: bOP larvae in the water-only arena with flow; Static: bOP larvae in water-only arena without flow) and avoidance assays (bOP larvae, left OP-intact (L) or right OP-intact (R) uOP larvae, and nOP larvae in arenas with cadaverine stream in the stimulus zone). f Upper panel: illustration of navigational strategies that may be adopted after encountering a noxious chemical. Lower panel: schematics of kinematic parameters that can be extracted. | | denotes absolute value. g Histograms of turn angle distributions of first (upper) and subsequent (lower) bouts (i.e., final Δθ). h First bout average angular velocity (i.e., |final Δθ|/Δt) upon entering the rightmost zone. i Swim bout frequency quantified from all rightmost zone entry-to-escape trajectories. j |Final Δθ| vs. bout number (line: mean; shadow: SEM) after rightmost zone entry. P value: two-sided Mann–Kendall trend test. b, d, e, i The parameters are plotted in log scales. In b, d, e, h and i Horizontal lines indicate the medians, 75 and 25 percentiles for each group. Shadows of the violin plots scale according to the probability density function. P values: Kruskal–Wallis test with Tukey's post hoc test. In a–e and g–j, numbers of assays, larvae, and rightmost zone border-crossing events: bOP (control): 6, 24, 211; bOP (avoidance): 8, 32, 251; uOP-L: 4, 15, 71; uOP-R: 4, 15, 155; nOP: 7, 23, 96; bOP Static: 3, 11, 283. Source data are provided as a Source Data file. Credit: Nature Communications (2023). DOI: 10.1038/s41467-023-35836-2 -
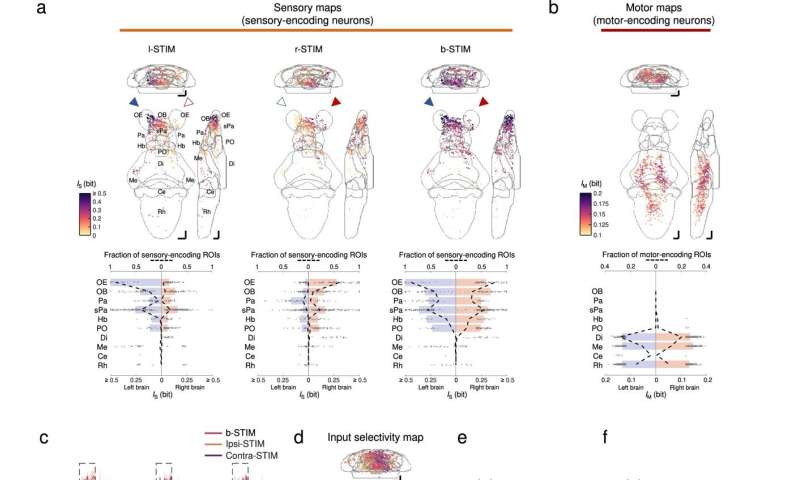
Brainwide neuronal activities evoked by chemosensory signals. a Upper panels: mean intensity projections (to coronal, transverse, and sagittal planes) of the mutual information between the calcium signals of regions-of-interest (ROIs) and stimulus profile of l-STIM (left panel), r-STIM (middle panel) or b-STIM (right panel) (IS), from an example larva. Solid triangles mark the corresponding OP(s) stimulated. Lower panels: corresponding brainwide IS distributions. Dashed lines indicate sensory-encoding ROI fractions. Bars represent the medians in regions with top six fractions of sensory-encoding ROIs with b-STIM. Abbreviations of brain regions: same as in Fig. 3e. b Upper panel: mean intensity projections of mutual information between the calcium signals of ROIs and tail flipping frequency (IM) from the larva in a. Lower panel: corresponding brainwide IM distribution. Dashed lines indicate motor-encoding ROIs fractions. Bars represent the medians in regions with top three fractions of motor-encoding ROIs. a, b The numbers of sensory-encoding and motor-encoding ROIs are 676 and 763, respectively. c Example trial-averaged responses to ipsilateral (ipsi-STIM, orange), contralateral (contra-STIM, violet), or bilateral (b-STIM, cherry) olfactory stimulation (n = 3 trials for each case) of individual ROIs from the designated brain regions with a range of ipsilateral-contralateral input selectivity (first number) and fraction of nonlinear information (FIs) (second number). Shadow shows SEMs. Dashed rectangle indicates stimulus window. Scale bars: 10 seconds (horizontal) and 0.5 normalized dF/F (vertical). Data from the same larva shown in a. d Mean intensity projection maps of ipsilateral(Ipsi)-contralateral(Contra) input selectivity of sensory-encoding ROIs. e Brainwide Ipsi-Contra input selectivity distributions of individual ROIs. f Regional means of Ipsi-Contra input selectivity. g Mean intensity projection maps of FIs. h Brainwide FIs distributions of individual ROIs. i Regional means of FIs. d–i Data are pooled across larvae (n = 4). The number of ROIs in d and e is 2301, and that in g and h is 1232. e, f, h, i The colors are coded accordingly to the color scale bars in d and g, respectively. e, h Horizontal lines: medians, 75 and 25 percentiles. Shadows of the violin plots scale according to the probability density function. f, i Each small dot representing one larva's value. Large dots, upper and lower limits of lines: medians, 75 and 25 percentiles, respectively. Scale bars in a, b, d, g 50 μm in Z-Brain atlas space. Source data are provided as a Source Data file. Credit: Nature Communications (2023). DOI: 10.1038/s41467-023-35836-2
Outlook
In this way, Samuel Sy and colleagues built on existing experimental advances established across the past decade within circuit neuroscience. They credit their research advances in part to newer methods that permit comprehensive behavioral monitoring and large-scale cellular-resolution of neuronal activities in diverse regions of the brain. The team developed an optofluidic method to assess chemosensory-mediated actions and brain-wide neural representations in larval zebrafish integrated with light-sheet fluorescence microscopy for whole brain imaging.
The microfluidics-based behavioral, chemical delivery and imaging principles demonstrated with Fish-on-Chips can be readily adapted to study chemosensory behaviors in several other organisms of similar size or smaller length scales such as bacterial and larvae of drosophila as well. While wild animals in the scale of interest can navigate more complex odor landscapes, the Fish-on-Chips navigation platform offers a precisely regulated optofluidic setup to comparatively assess the neural representation of individual odors and detect the ensuing behavior of organisms. This experimental device will allow the team to gain deeper insights into the principles of sensory information processing.
More information: Sy et al, An optofluidic platform for interrogating chemosensory behavior and brainwide neural representation in larval zebrafish, Nature Communications (2023). DOI: 10.1038/s41467-023-35836-2
Fan Yang et al, Fish-on-a-chip: microfluidics for zebrafish research, Lab on a Chip (2016). DOI: 10.1039/C6LC00044D
Journal information: Nature Communications , Lab on a Chip
© 2023 Science X Network





















