Genomes OnLine Database introduces new features
Since its launch 25 years ago, the Genomes OnLine Database (GOLD) has matured from six projects on a spreadsheet into a flagship genomic metadata repository, making curated microbiome metadata that follows community standards freely available, and enabling large-scale comparative genomics analysis initiatives.
GOLD not only curates sequencing projects carried out at the U.S. Department of Energy (DOE) Joint Genome Institute (JGI), a DOE Office of Science User Facility located at Lawrence Berkeley National Laboratory (Berkeley Lab), but also those imported from public repositories, as well as project data entered by external users.
The Impact
One of the significant challenges in maximizing genomic data usage is having the appropriate contextual information, or metadata, stored with them. GOLD provides manually curated metadata resources for organisms and environmental samples. The database aggregates relevant metadata from various sources and applies a standardized labeling scheme to better define the genomic information and ecosystems.
These curated data are accessible on JGI data portals including the Integrated Microbial Genomes & Microbiomes (IMG/M) system, enhancing genome annotations and promoting comparative genome analyses. GOLD aims to follow FAIR data principles to ensure digital assets are findable, accessible, interoperable and reusable.
Summary
GOLD takes samples processed at the JGI, entered by external users and those imported from the public repositories and curates the information by applying community-developed standards. GOLD also supports a broad range of activities from proposal intake to publication. Information is also obtained and cross-checked with public resources including NCBI Taxonomy and various culture collections like the American Type Culture Collection and the Leibniz Institute DSMZ. GOLD applies its hallmark standardized naming for all the environmental samples and is the only resource in the world with nearly 200,000 curated environmental samples with canonical names.
Since its 1997 launch, active GOLD users have spurred its growth and the continuing development of new components and capabilities. The most recent improvements include new features like a public API and ecosystem landing page, as well as the growth of different entities, further outlined in the journal Nucleic Acids Research.
One of GOLD's strengths has been implementing metadata standards across all entities in the system. The JGI's Genomic Standards Group, which manages GOLD, communicates personally with submitters to resolve any inconsistencies. While in the past, GOLD relied on free text fields, it now leverages controlled vocabularies with unit-based fields like depth and elevation recorded in fixed units of meters, temperature in centigrade, and more.
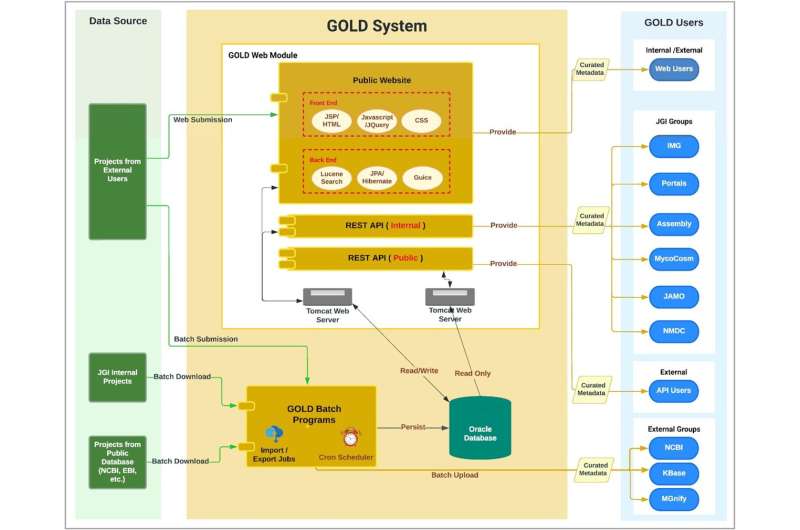
The four level project organization system implemented in GOLD consist of: Studies, Biosample/Organisms, Sequencing Projects and Analysis Projects. Each of these entities are curated with a wide range of metadata. GOLD applies a five-level ecosystem classification to all environmental samples and organisms whose isolation information is available—enabling metadata-driven scientific discoveries. GOLD's public Application Programming Interface allows users to access curated metadata programmatically in a secure and reliable manner.
The JGI team managing GOLD already works closely with two other DOE-funded projects—the National Microbiome Data Collaborative (NMDC) and the DOE Systems Biology Knowledgebase (KBase)—and aims to extend these collaborations in metadata curation, establishing metadata standards and sample metadata exchange.
More information: Supratim Mukherjee et al, Twenty-five years of Genomes OnLine Database (GOLD): data updates and new features in v.9, Nucleic Acids Research (2022). DOI: 10.1093/nar/gkac974
Journal information: Nucleic Acids Research
Provided by DOE/Joint Genome Institute