Database of DNA viruses and retroviruses debuts on IMG platform
In a series of four articles published in the Database issue of the Nucleic Acids Research journal, DOE JGI researchers report on the latest updates to several publicly accessible databases and computational tools that benefit the global community of microbial researchers. One report focuses on a new database dedicated global viral diversity.
Microbes play key roles in maintaining the planet's biogeochemical cycles. Viruses, thought to outnumber microbes by 10-fold, exert major influences on microbial survival and community interactions. Advances in sequencing technologies have generated vast amounts of data about these viruses, requiring tools to manage and interpret the information. These updates focus on database analytical tools for microbial genomics and viruses relevant to DOE missions in bioenergy and environment.
Providing high-quality, publicly accessible sequence data goes hand-in-hand with developing and maintaining the databases and tools that the research community can harness to help answer scientific questions. In the Database issue of the journal Nucleic Acids Research, which will be released January 1, 2017, researchers at the U.S. Department of Energy Joint Genome Institute (DOE JGI), a national user facility, describe a database called IMG/VR (https://img.jgi.doe.gov/vr/), IMG/VR is the largest such publicly available database, with 3,908 isolate reference DNA viruses and 264 413 computationally identified viral contigs from >6000 ecologically diverse metagenomic samples.
A comprehensive computational platform integrating all these sequences with associated metadata and analytical tools accompanies IMG/VR, which follows on the heels of a recent DOE JGI viral diversity study report in Nature. Additional articles in the same issue describe updates to several publicly accessible, interactive databases since the last set of reports published in 2014. For example, as of July 2016, there were 47,516 archaeal, bacterial and eukaryotic genomes in the Integrated Microbial Genomes with Microbiome Samples (IMG/M: https://img.jgi.doe.gov/m/) system, with researchers noting that number "represents an over 300% increase since September 2013." IMG/M contains: annotated DNA and RNA sequence data of archaeal, bacterial, eukaryotic and viral genomes from cultured organisms; single cell genomes (SCG) and genomes from metagenomes from uncultured archaea, bacteria and viruses; and, metagenomes from environmental, host associated and engineered microbiome samples.
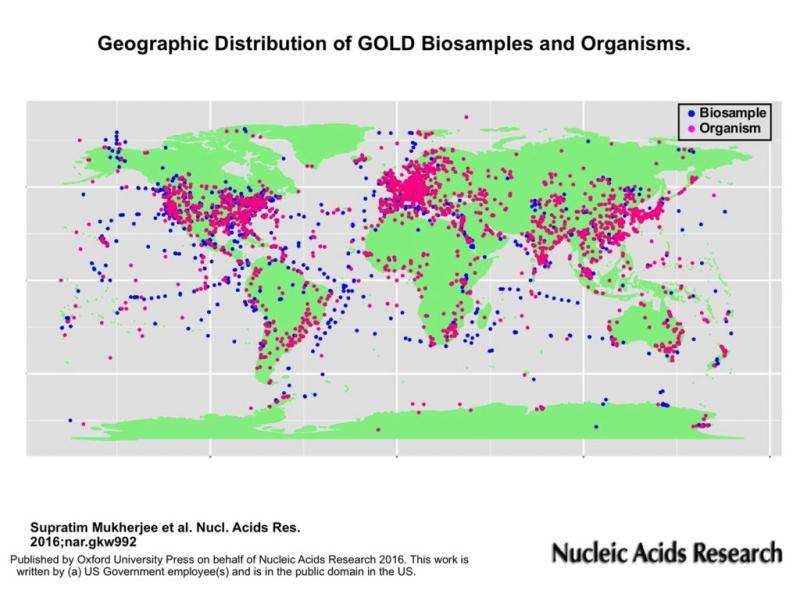
Another paper concerns the Genomes Online Database (GOLD: https://gold.jgi.doe.gov), a manually curated data management system that catalogs sequencing projects with associated metadata from around the world. In the current version of GOLD (v.6), all projects are organized based on a four level classification system in the form of a Study, Organism (for isolates) or Biosample (for environmental samples), Sequencing Project and Analysis Project. A fourth paper focuses on the Integrated Microbial Genomes Atlas of Biosynthetic gene Clusters (IMG-ABC: https://img.jgi.doe.gov/abc/). Launched in 2015, IMG-ABC allows researchers to search for biosynthetic gene clusters and secondary metabolites and their latest update now incorporates ClusterScout, a tool for targeted identification of custom biosynthetic gene clusters across several thousand isolate microbial genomes, and a new search capability.
More information: Chen IA et al. IMG/M: integrated genome and metagenome comparative data analysis system. Nucleic Acids Res. 2016 Oct 13. pii: gkw929. [Epub ahead of print]
Mukherjee S et al. Genomes OnLine Database (GOLD) v.6: data updates and feature enhancements. Nucleic Acids Res. 2016 Oct 27. pii: gkw992.
Paez-Espino D et al. IMG/VR: a database of cultured and uncultured DNA Viruses and retroviruses. Nucleic Acids Res. 2016 Oct 30. pii: gkw1030. [Epub ahead of print]
Hadjithomas M et al. IMG-ABC: new features for bacterial secondary metabolism analysis and targeted biosynthetic gene cluster discovery in thousands of microbial genomes. Nucleic Acids Res. 2016 Nov 29. pii: gkw1103. [Epub ahead of print]
Journal information: Nucleic Acids Research
Provided by DOE/Joint Genome Institute