Better microbial genome binning with metaBAT
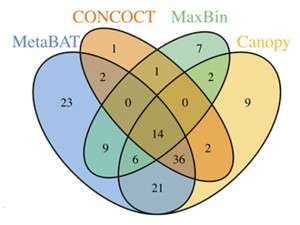
DOE JGI researchers have developed an automated tool called MetaBAT that automatically groups large genomic fragments assembled from metagenome sequences to reconstruct single microbial genomes.
The ability to accurately and efficiently reconstruct individual microbial genomes from large and complex metagenome datasets allows researchers to understand how they each interact with each other and influence global cycles. Tools such as MetaBAT enable researchers to better appreciate the data generated by high-throughput metagenome sequencing, which allows researchers to study microbial communities without needing to cultivate them.
If a few grams of soil can hold a multitude of microbes interacting with each other to influence the global carbon cycle, imagine how many microbes in a cow's rumen are contributing to efforts in breaking down the plant mass for nutrients, information that could prove useful in developing sustainable alternative fuels from plants. Technological advances have allowed researchers to utilize high-throughput metagenome shotgun sequencing to study microbial communities without needing to cultivate these organisms. However, researchers are still working on ways to efficiently and accurately assemble individual microbial genomes from these large-scale datasets to learn more about each one's specific contributions to maintaining the global cycles.
Most of the current approaches to "bin" or group large genomic fragments from metagenome datasets in order to reconstruct individual genomes have limitations. Some of them rely on known genomes as references, an approach that does not work well on environmental samples where many microbes do not have closely related species with known genomes. Many of them contain manual steps and do not scale up to handle large metagenomic datasets.
In a paper published August 27, 2015 in PeerJ, researchers from the U.S. Department of Energy Joint Genome Institute (DOE JGI), a DOE Office of Science User Facility, offer an automated metagenome binning software tool that resolves these obstacles. The paper describing MetaBAT (for Metagenome Binning with Abundance and Tetra-nucleotide frequencies) is among the 10 articles selected as representing "some of the most noteworthy genomics research which PeerJ has published in the 2 years prior to October 2015."
The team evaluated MetaBAT using both synthetic and real-world metagenome datasets, comparing the number of genome bins accurately recovered with the tool against those found by other binning methods. They found that MetaBAT recovered "many [genomes] missed by alternative tools." More importantly, they reported that MetaBAT was computationally efficient; the software identified 340 bins from the synthetic metagenome dataset with 200,000 fragments in only 14 minutes and using only 3.9 GB of RAM while other methods took 20 to 104 hours and used 9.5GB to 38GB of RAM for the same data.
MetaBAT is an open source software tool available at bitbucket.org/berkeleylab/metabat along with a software manual. The tool has been used by researchers in a Nature study on applying omics to study microbial communities in the permafrost, as well as in a Science study reporting the reconstruction of complete genomes of deep groundwater methanogens from an archaeal phylum that had not been previously associated with methane metabolism.
More information: Dongwan D. Kang et al. MetaBAT, an efficient tool for accurately reconstructing single genomes from complex microbial communities, PeerJ (2015). DOI: 10.7717/peerj.1165
Journal information: PeerJ
Provided by DOE/Joint Genome Institute