A rallying call for microbiome science national data management
Massive amounts of data require infrastructure to manage and store the information in a manner than can be easily accessed for use. While technologies have scaled to allow researchers to sequence and annotate communities of microorganisms within an environment,(its "microbiome"), on an ever-increasing scale, the data management aspect has not been developed in parallel.
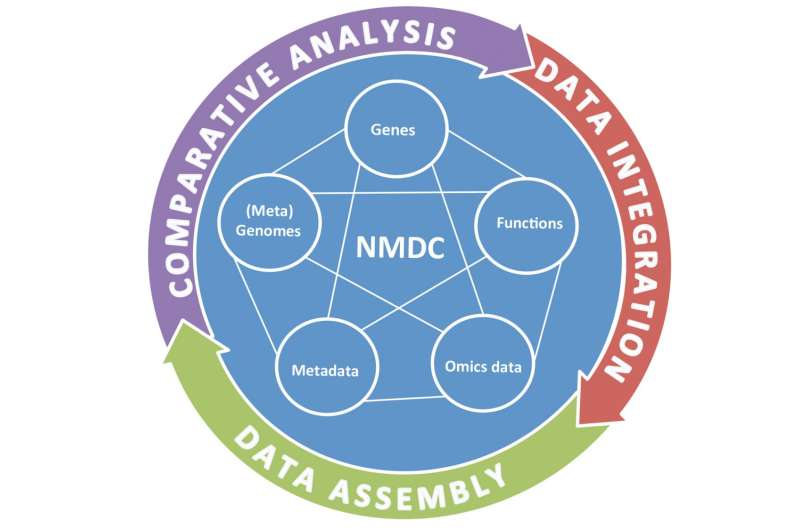
In a paper published online May 16, 2016 in Trends in Microbiology, researchers from the U.S. Department of Energy Joint Genome Institute (DOE JGI), a DOE Office of Science User Facility, call for the formation of a National Microbiome Data Center to efficiently manage the datasets accumulated globally. By integrating and harnessing all available microbiome data and metadata, researchers could conduct larger-scale comparative analyses in order to address global challenges related to energy, environment, health and agriculture.
"The time is ripe to embark on the greatest endeavor to understand Earth's microbiome," said Nikos Kyrpides, DOE JGI Prokaryote Super Program head and the study's first author. "Biological sequence data should be considered an instrumental tool for the study of biology systems, analogous to the telescope for astronomy and the particle accelerator for high-energy physics."
A Complement to the National Microbiome Initiative
The timely publication complements the White House's launch of a National Microbiome Initiative focused on comparing microbial communities across ecosystems to identify the "organizing principles" that shape all microbiomes. A national microbiome data center, the team wrote, would "organize, process, and serve all available environmental genomic data."
Kyrpides and his colleagues identified three bottlenecks in microbiome research associated with short-sightedness: lack of a grand vision to move beyond "single-use" microbiome datasets to a more cohesive collection; lack of interagency funding models; and, limited international data standards that hinder the global research community's ability to efficiently conduct comparative analyses. Several large data management systems already exist to help, including the Integrated Microbial Genomes (IMG) system and the Genomes OnLine Database (GOLD) system run by DOE JGI scientists. These resources allow researchers to access and analyze publicly available assembled microbial and microbiome data and metadata, respectively. In addition, the DOE JGI has partnered with the National Energy Research Scientific Computing Center (NERSC) to operate in a high performance computing environment and support the growing community demand.
A Grand Vision as Microbiome Research Scales
"There is a profound lack of a grand vision in appropriate funding to support the extraction of knowledge from big data (i.e., across studies)," Kyrpides said. "Furthermore, the reference data needed to contextualize the myriad microbiome samples is sorely lacking. These data are fundamental for interpretation of how microbiomes function, and how they interact within the environments and hosts they inhabit. Systematic decoding of microbes and their environments to fill in the gaps in our databases is a key step towards hypothesis-driven science and enabling a better understanding of microbial life."
The Department of Energy has a tradition of taking on massive projects—from the first particle accelerator to its role in initiating the Human Genome Project, and the DOE JGI is no stranger to microbiome research, reporting the first genomic characterization of a microbial community back in 2004. Over the past decade, microbiome research has grown in scale, tackling projects such as termite hindgut, cow rumen, the Gulf of Mexico oil-eating microbiome, prairie soils and permafrost. Through the Community Science Program, the largest dataset focused on oxygen minimum zones and what has been described as the "only systematically and quantitatively prepared dataset available" for the viral ecology community were developed in collaboration with the DOE JGI.
"At the dawn of the third decade of microbial genomics, and well into the information age, the establishment of a national microbiome data center can pave the way to understanding the Earth's microbiome," Kyrpides said.
More information: Nikos C. Kyrpides et al, Microbiome Data Science: Understanding Our Microbial Planet, Trends in Microbiology (2016). DOI: 10.1016/j.tim.2016.02.011
Provided by DOE/Joint Genome Institute