Learning-based strategy to predict gene expression and identify regulatory sequences
The research team led by Prof. Guo Guoji and Prof. Han Xiaoping at the Zhejiang University School of Medicine have published an article titled "Deep learning of cross-species single cell landscapes identifies conserved regulatory programs underlying cell types" in the journal Nature Genetics on October 13.
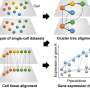
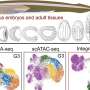
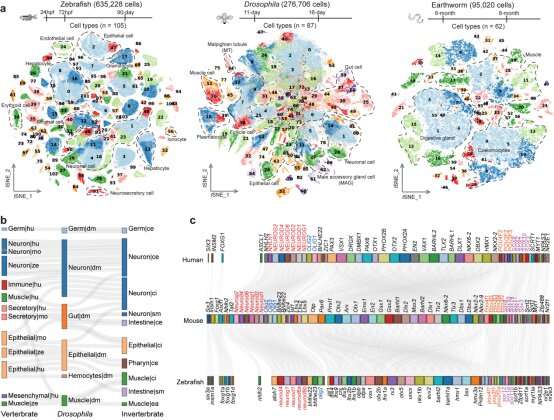
In their study, the researchers employed their independently developed Microwell-seq to construct organism-wide cell landscapes for zebrafish, Drosophila and earthworms using a whole-body strategy that could eliminate tissue-specific batch effects. Specifically, they analyzed 635,228 single cells from zebrafish, 276,706 single cells from Drosophila, and 95,020 single cells from earthworms. Together with other five cell landscapes, they analyzed a total of eight representative metazoan species to explore conserved genetic regulation in vertebrates and invertebrates.
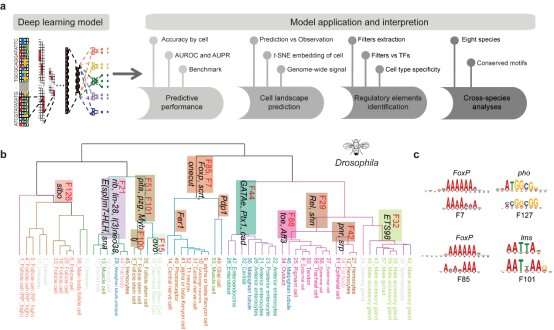
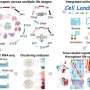
Most important of all, they developed a deep-learning-based framework, Nvwa (the name of a mother god in ancient Chinese legend), to predict gene expression solely from the DNA sequence at the single cell level. Notably, Nvwa can accurately predict gene expression in virtually all studied species. By extracting the deep-learning-based motifs from each first-layer convolution filter, they interpreted the cell-type-specific sequence rules and identified conserved regulatory programs across species.
It is the first time that an integrated model has been created for cross-species transcriptomic landscapes. This study provides a valuable resource and offers a new approach to study regulatory grammar in diverse biological systems.
More information: Jiaqi Li et al, Deep learning of cross-species single-cell landscapes identifies conserved regulatory programs underlying cell types, Nature Genetics (2022). DOI: 10.1038/s41588-022-01197-7
Journal information: Nature Genetics
Provided by Zhejiang University