Breakthrough in CRISPR research may lead to more effective and safer gene editing
10 years ago we saw a breakthrough in modern biology.
An American scientist discovered that manipulation of the Cas9 protein resulted in a gene technology worthy of a sci-fi film: CRISPR.
Think of it as a pair of molecular scissors capable of cutting and editing the DNA of humans, animals, plants, bacteria and viruses.
The potential is huge and covers anything from deleting hereditary diseases to producing crops able to withstand climate change.
However, like any other new technology, CRISPR has had its challenges. One of the main challenges has been to make the technology as effective as possible and to make sure the scissors only cut where we want them to.
'We have described new mechanisms behind CRISPR'
Two new studies from the University of Copenhagen conducted together with researchers from Aarhus University can help solve these problems.
"We have described new mechanisms behind CRISPR," says Professor of Bioinformatics Jan Gorodkin from the Department of Veterinary and Animal Sciences.
"We are now able to explain why some off-targets, which is unintended cuts elsewhere in the genome, are more effective than on-targets, which is the cut at the intended place. We have also learned how different DNA sequences around the on-target can impact how efficiently the Cas9 protein cuts the DNA. Hopefully, this knowledge will pave the way for more effective and safer use of CRISPR."
So how does CRISPR work? First, a scientist will design a piece of synthetic RNA called the guide RNA. This is then attached to the Cas9 protein which will perform the task of cutting the DNA. The guide RNA scouts for the matching DNA section. Once the guide RNA has found the right spot, Cas9 will cut the DNA string. Now the scientist is able to insert any synthetic piece of DNA into the vacated spot.
If Cas9 and the guide RNA hit the target, the scientists refer to it as being on target; if they hit another place, they are off target.
Today, CRISPR is in the context of medicine mainly used to study how genes and drugs work in the laboratory and is still not widely used in human treatment. However, in the long term, the idea is to use CRISPR in the treatment of certain genetic diseases.
Mystery of effective off-targets solved
In one of the two new studies, the researchers sought to determine the best way for the guide RNA to attach to the DNA, making the cut as effective as possible—because if the cut is not sufficiently effective, the scientists will not be able to edit the DNA.
"We already know that CRISPR does not really work when the bond between the guide RNA and the DNA is too weak. Now we have learned that too strong a bond is problematic too," says Gorodkin. "In both cases, the gene scissors are too weak and ineffective."
The researchers then identified an interval where the bond between the guide RNA and the DNA is neither too strong nor too weak, but just right, resulting in scissors with perfect sharpness.
"Interestingly, this observation can also be used to explain why some off-targets show stronger CRISPR activity than their intended on-targets—that is, why the scissors are sharper for some off-targets than for the on-targets," Gorodkin explains.
"This is because too strong on-targets do not fall within the right binding energy interval. But if you remove some of the energy from these strong bonds, which is what is happening at the off-target sites, you are able to get into the interval right, resulting in a more powerful effect and thus a shaper scissor at the off-targets."
The study has also identified the optimal position of the Cas9 protein for achieving the most effective cut.
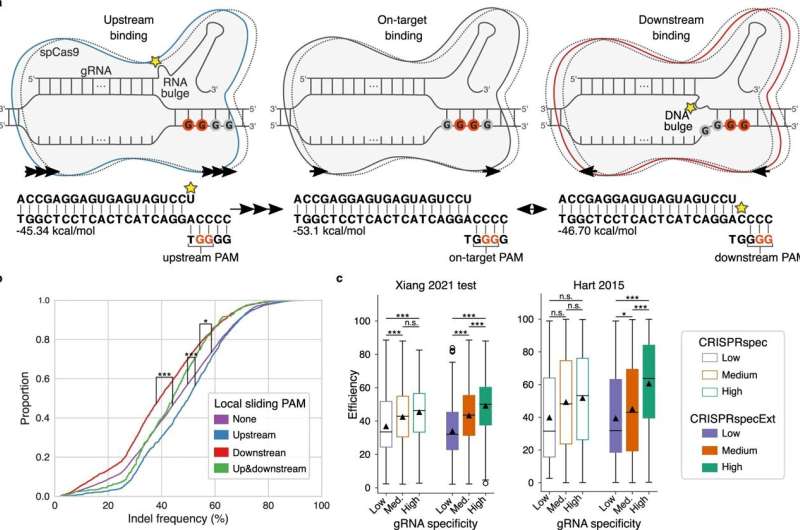
Before Cas9 is able to cut the DNA, the protein must bind to a specific part of the DNA string. DNA consists of four different nucleotides: A, C, G and T, and Cas9 can only bind to a sequence with two consecutive G nucleotides.
Now the researchers have identified the effect on Cas9 of multiple consecutive G nucleotides—a situation where it is hard to hit the target because every two consecutive G's compete for binding with Cas9.
"When there are several G's 'upstream', that is, before the sequence that Cas9 was intended to bind to, the cut will be more effective. But when there are several G's 'downstream', that is, after the intended sequence for Cas9 to bind to, the cut is less efficient," explains Postdoc Giulia Corsi.
Corsi hopes the new knowledge into how CRISPR works will make it easier to identify the correct position of Cas9 in the future. This should also help minimize the number of potential side effects.
"We would like to be able to predict the cut, improve target editing and eliminate off-targets, which complicate the development of new drugs by requiring lots of resources and can result in side effects that occur when you cut the wrong gene," says Corsi.
Off-targets can be harmful—and they are under-researched
The second study focuses on off-targets. Here the researchers developed a method for measuring the effectivity of off-targets.
To quality-control a CRISPR experiment, scientists will usually select a smaller number of computer-predicted off-targets for testing. Using the new technology, however, they will be able to test a much larger number of off-targets, and this is expected to speed up the development of new drugs with fewer side effects.
Using the new method, the researchers tested 8,000 potential off-targets for 110 CRISPR guide RNAs in the process of being translated into human medicine. They found that around 10 percent of the 8,000 potential off-targets were in fact off-targets.
"We found far more off-targets than we would have been able to using existing methods," Gorodkin explains.
Furthermore, 37 of these off-targets are located in cancer related genes, which increases the risk that developing drugs will become harder or even impossible. In addition, unintended cuts in these genes may even lead to cancer as a possible side-effect.
"Researchers need to be able to identify such off-targets and select other guide-RNAs which do not have these or any other critical off-target," says Gorodkin.
Great need for more research into off-targets
According to Gorodkin, this shows that there is a great need for more research into off-targets.
"I would argue—and some may disagree—that off-targets are extremely under-researched. My impression is that existing studies on gene editing often lack the complete tools and analyses required to show that there are no off-target effects in their studies," he says.
According to Jan Gorodkin, the new method will have great impact in the future to better check studies for off-targets.
"In the past 10 years, we have taken a big step towards being able to edit the genome. Now we are in the process of making our methods better, safer and more effective. The latter also supports the green transition, as genome modifications, e.g. of cells used in production, can lead to more cost-effective utilization of resources."
The two studies are published in Nature Communications.
More information: Giulia I. Corsi et al, CRISPR/Cas9 gRNA activity depends on free energy changes and on the target PAM context, Nature Communications (2022). DOI: 10.1038/s41467-022-30515-0
Xiaoguang Pan et al, Massively targeted evaluation of therapeutic CRISPR off-targets in cells, Nature Communications (2022). DOI: 10.1038/s41467-022-31543-6
Journal information: Nature Communications
Provided by University of Copenhagen