Study develops new way of identifying cancer cells
A new method of separating cancer cells from non-cancer cells has been developed by researchers at the Wellcome Sanger Institute, in a boost for those working to better understand cancer biology using single-cell mRNA sequencing.
The study, published today in Communications Biology, improves on existing methods to identify which cells in a sample are cancerous and provides crucial data on the microenvironment of tumors. The software is openly available for researchers around the world to apply to their own data, advancing the effectiveness of single-cell sequencing to understand cancer.
Single cell mRNA analysis of cancer cells is one of the leading edge techniques being used to better understand cancer biology. The data generated can be used to try to disrupt cancers with drugs or work out how cancers arise in the first place.
A fundamental step in this process is separating cancer and non-cancer cells, but this isn't always an easy task. As well as the many types of cancer, there will also be molecular differences between cancer cells of the same type within a single tumor.
Currently, the best method of doing this is to measure the average gene expression of cells in the sample, with higher or lower expression used to distinguish cancer cells from healthy cells. But this method can lead to false conclusions.
In this new study, researchers at the Wellcome Sanger Institute performed whole genome sequencing and single-cell mRNA sequencing on samples collected by Great Ormond Street Hospital (GOSH).
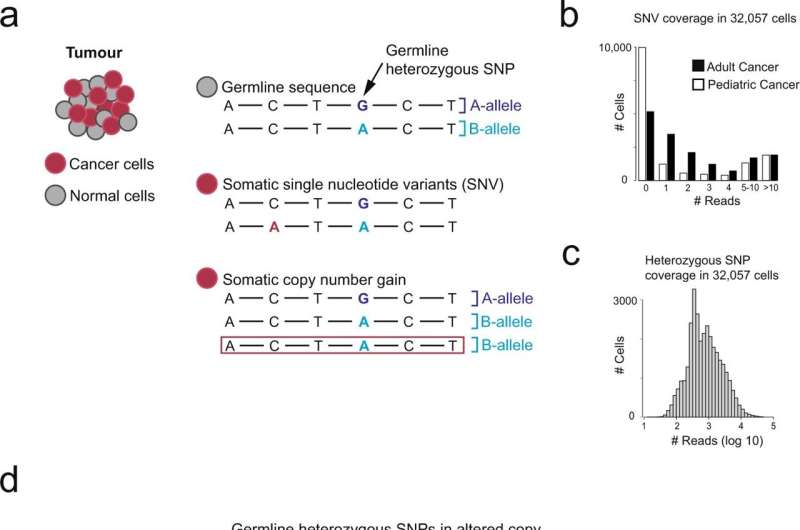
By locating imbalances of alleles in these data, which indicate copy number changes in the genome, the team was able to identify cancer cells more reliably than with previous methods. This approach will primarily be useful for validating new cancer cell types and better understanding the microenvironment of tumor tissue.
"Being able to know how the transcriptome is different in cells with aberrant genomes, such as those found in cancers, is valuable knowledge and will expand the questions that we can answer using single-cell sequencing," says Dr. Matt Young.
The method, named alleleIntegrator, is available as a software package for researchers across the world to use.
More information: Mi K. Trinh et al, Precise identification of cancer cells from allelic imbalances in single cell transcriptomes, Communications Biology (2022). DOI: 10.1038/s42003-022-03808-9. www.nature.com/articles/s42003-022-03808-9
Journal information: Communications Biology
Provided by Wellcome Trust Sanger Institute