Single-cell RNA sequencing to study salmonella infection
In May 2022, the U.S. Food and Drug Administration (FDA) recalled Jif peanut butter due to potential salmonella contamination. In the last 5 years alone, there have been ~35 food-related salmonella outbreaks. Salmonella enterica is a gram-negative pathogen that typically invades the intestines to cause disease. People who get salmonella infection experience symptoms with varying degrees of severity—many have diarrhea and fever, while some have no symptoms at all. So, what dictates the disease outcome during salmonellosis? There can be myriad factors at play, like differences in the strains of salmonella that cause infections, or variations in the genetic make-up or immune responses of individuals. In other words, not all cells (be they the host or the pathogen) are the same or have similar physiology. These factors can result in a wide clinical spectrum of salmonella infection.
Resolving single cells by RNA sequencing
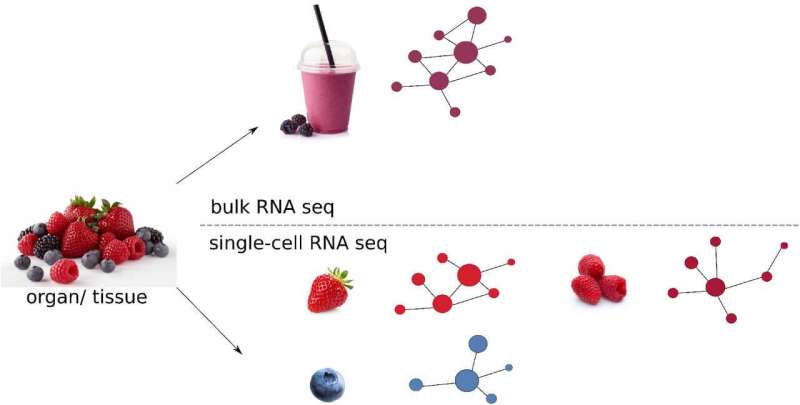
Scientists are now trying to dissect this heterogeneity in host-salmonella interactions by investigating the behavior of single cells at an unprecedented resolution. "This is very challenging to study as bacterial numbers are minute during different stages of salmonella infection. So, we need sensitive tools to probe them," said Prof. Roi Avraham, Principal Investigator in the Department of Immunology and Regenerative Biology at the Weizmann Institute of Science, during a session at ASM Microbe 2022. One technique that is sensitive enough to dissect these complex interactions is RNA sequencing (RNA-seq). Traditional bulk RNA-seq reveals the average transcriptome—expression levels of different genes in cells—by quantifying messenger RNA, or mRNA, in a given sample.
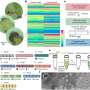
In the past decade, scientists have unleashed the potential of single-cell RNA sequencing (scRNA-seq) to investigate host-microbe interactions. scRNA-seq provides information about the transcriptome in single cells, offering a high spatiotemporal view of the infection landscape. An analogy that is often mentioned in reference to single-cell techniques is that of a fruit smoothie—while bulk measurements can provide information about the overall characteristics of the fruit smoothie, single-cell technologies provide information about individual pieces of strawberry, banana, apple and so forth.
By understanding which genes are upregulated or downregulated in individual cells, one can get clues as to why different cells behave the way they do. The development of targeted antimicrobial therapies for treating Salmonella infections with different clinical outcomes remains a primary goal of such advanced understanding.
Cellular heterogeneity during salmonella infection
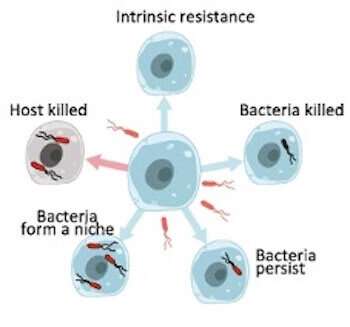
When a pathogen infects the host cell, the pathogen activates a virulence program to cause infection, whereas the host activates a defensive immune response to combat the invader. A lot of factors can influence what will happen in this tug-of-war. For instance, when and where does the pathogen trigger its virulence program, or what genes are turned on or off in the infected cells at a given time during infection?
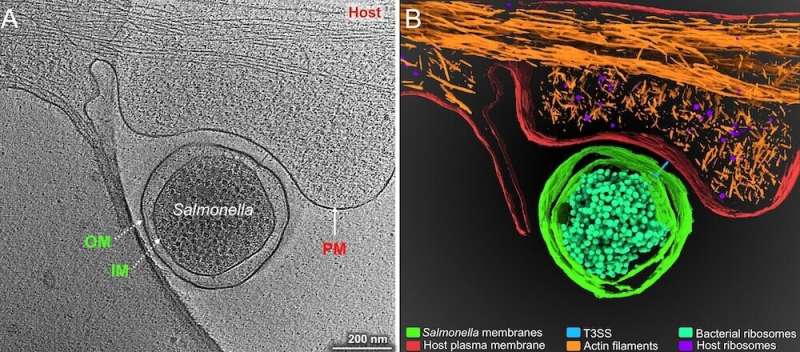
Salmonella enterica serovar Typhimurium or Typhimurium (S. Typhi) invades the host intestinal cells and begins its intracellular life cycle inside phagocytes like macrophages. In doing so, the bacterium escapes the host immune responses and later spreads throughout the body via the bloodstream.
It is known that S. Typhi has heterogeneous replication rates inside both macrophages and tissues. One subpopulation of S. Typhi grows fast and produces proteins that hijack the host cellular machinery to favor the survival of the pathogen. These virulence proteins are called effectors and are produced by 2 type III secretion systems (T3SSs) that are located on Salmonella Pathogenicity Island (SPI) 1 and 2. Meanwhile, a second subpopulation of S. Typhi remains as non-replicating cells called persisters. Persisters can display further heterogeneity, depending on their metabolic activities. They are also resistant to many antibiotics and, hence, can serve as a reservoir for chronic infections.
S. Typhi further displays heterogeneity during infection in response to fluctuating host immune responses, which could potentially reprogram the host cellular machinery into distinct subpopulations. "The pathogen itself influences the host immunity and is influenced by the immunity. It has to balance its replication, virulence activities and manipulation of host immunity," said Avraham.
scRNA-seq of macrophages infected with S. Typhi revealed that actively dividing salmonella are predominantly found in M2-like macrophages that are anti-inflammatory and support bacterial growth. On the other hand, non-replicating salmonella are predominantly found in M1-like macrophages that are pro-inflammatory and arrest bacterial growth. Non-replicating salmonella persisters make up a smaller subpopulation compared to their fast-growing counterpart, but they are clinically significant as they are resistant to antimicrobials.
When antibiotics are removed, persisters can cause disease relapse. The reason for this is that even though they do not grow and divide, they are still metabolically active and able to reprogram macrophages via SPI-2 virulence effectors. This suggests that variations in gene expression in host macrophages can create different cellular environments that can affect the pathogen in distinct ways—either by allowing salmonella to escape host immunity and survive under stealth, or by allowing them to escape host antimicrobial defenses and grow.
Dual RNA-seq studies for host-salmonella
Most of the studies thus far have applied scRNA-seq to profile either the host or the pathogen, but researchers are now working to evaluate both host and pathogen cells simultaneously in an approach called dual RNA-seq. Application of bulk dual RNA-seq in salmonella-infected host cells revealed that a bacterial small RNA called PinT manipulates the expression of many host and bacterial genes. PinT upregulates genes involved in inflammatory signaling in the intestines of the host, a reaction salmonella depends on to compete with gut microbes and avoid being killed by the host. On the bacterial side, PinT downregulates the expression of certain SPI-1 and SPI-2 virulence effectors that allow the bacteria to replicate intracellularly.
Researchers are now building a single-cell version of dual RNA sequencing (scDual-seq) to study host-salmonella interactions. During ASM Microbe 2022, Avraham introduced a technique developed in his lab called scPAIR-seq to analyze pathogen-specific immune responses in the context of S. Typhi infection. This technique enables researchers to study the impact of multiple bacterial mutants on host cells, simultaneously and at single-cell resolution.
In this work, researchers infected macrophages with a library of uniquely barcoded bacteria that have mutations in SPI-2 virulence effectors to investigate mutant-specific changes to the host transcriptome. They found that among virulence effectors, SifA had the most significant impact on the host transcriptome. SifA helps salmonella to establish a protective niche inside vacuoles in macrophages. In the absence of SifA, S. Typhi is present in the cytoplasm instead of vacuoles and reprograms infected macrophages to an anti-inflammatory M2 state. This allows the bacteria to proliferate in the cytosol, suggesting an alternative route of infection. The experimental and computational pipeline developed in this study can be used to study the effect of multiple effectors simultaneously in other pathogenic bacteria, as well to provide a global and holistic view of effectors on host immune pathways.
Applications of scRNA-seq
RNA-seq, and in particular scRNA-seq, is becoming an increasingly popular choice to analyze changes in gene expression during host-pathogen interactions. These methods have revealed new insights into S. Typhi infections and provided clues about heterogeneity in both host cells and bacteria. Such studies can reveal molecular details of how immune cells eliminate pathogens and about the spatiotemporal targeting efficacy of different antimicrobial therapies.
Efforts are underway to build cellular atlases in various model organisms to reveal different cell types and states. Immunological Genome Project is one such effort that is creating gene expression profiles for all mouse immune cell types and is bound to play a pivotal role in host-pathogen interactions. This atlas can be used to understand which genes are modulated in different immune cell types during bacterial infections. Using such resources, one can begin to answer questions, such as why do bacteria infect or proliferate in some cells and not others, which host immune cells are more susceptible to infection and why.
Next stop: The clinic?
Scientists are also making efforts to translate RNA-seq findings from the laboratory to the clinic. In one such attempt, researchers applied an algorithm based on scRNA-seq data to predict outcomes of salmonella infection in healthy individuals ex vivo using their blood samples. Individuals with different immune cell types—either wild type or with a polymorphism in the TLR10 gene that regulates cytokine responses—demonstrate different infection phenotypes. Future work to expand cohort sizes and predict clinical outcomes of infectious diseases by analyzing heterogeneity at the cellular level can help develop personalized therapies for individual patients.
Provided by Weizmann Institute of Science