Understanding how motor proteins shape our cells
Understanding the busy networks inside our cells can help researchers develop new cancer treatments and prevent dangerous fungal infections.
With the help of the Canadian Light Source (CLS) at the University of Saskatchewan, a research team led by John Allingham from Queen's University and Hernando Sosa from the Albert Einstein College of Medicine has shed light on a protein that regulates the intricate microscopic networks that give cells their shape and helps ship important molecules to diverse locations.
Using the CMCF beamline at the CLS and the cryo-EM facility at the Simons Electron Microscopy Center (SEMC) at the New York Structural Biology Center, the team found the missing pieces of an important puzzle.
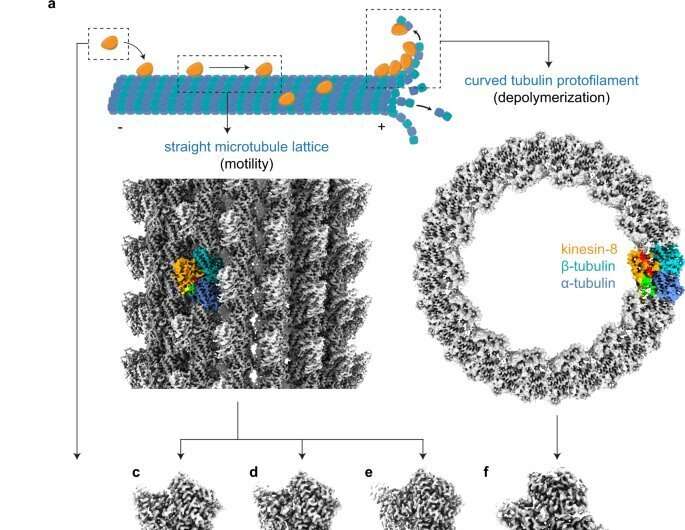
In their published work, they are the first group to clearly describe the mechanism of action of a tiny motor protein called Kinesin-8 that enables it to control the structures of microtubule fiber networks inside the cell.
"Our recent paper in Nature Communications, co-first authored by Byron Hunter and Matthieu Benoit, shows how this specific type of kinesin motor protein has developed the ability to use microtubules as tracks for movement, guiding transport of cargo within the cell," said Dr. John Allingham, a professor at the Queen's School of Medicine, "in addition to being able to disassemble these tracks, controlling their length and location in cells."
The Kinesin-8 proteins ensure that a cell's cargo is in the right place during cellular division and help to regulate cellular networks, making sure the microtubules do not grow too long.
This research provides an important strategy for cancer treatment. The team is hopeful that targeting the Kinesin-8 proteins in cancer cells could contribute to anti-cancer treatments.
This strategy could also be used to develop a therapy for pathogenic fungal infections that threaten people with compromised immune systems.
Allingham said the CLS provides an invaluable training environment for his students, including Ph.D. candidate Byron Hunter who collected the CLS data for their recent work.
"The CLS platform was hugely valuable," said Hunter. "The increase in the quality of data was enormous. We were able to screen a huge number of different crystal samples in a relatively brief period of time."
More information: Byron Hunter et al, Kinesin-8-specific loop-2 controls the dual activities of the motor domain according to tubulin protofilament shape, Nature Communications (2022). DOI: 10.1038/s41467-022-31794-3
Journal information: Nature Communications
Provided by Canadian Light Source