Researchers uncover molecular basis of antigen processing for cancer target MR1
In a new study published in Nature Chemical Biology, researchers at Children's Hospital of Philadelphia (CHOP) have revealed the molecular mechanics of a cell-surface molecule that is a potential immunotherapy target. Using sophisticated techniques, including nuclear magnetic resonance (NMR) spectroscopy, the researchers demonstrated the importance of so-called chaperone molecules in stabilizing this target, known as MHC-I-related protein 1 (MR1).
Major histocompatibility complex class I (MHC-I) molecules live on the surface of nearly all cells and present peptide antigens to the immune system, so that immune cells can detect when a foreign antigen is present, such as those from bacteria, viruses, or even developing tumors. In a similar process, the MHC-I-related protein 1 (MR1) presents smaller metabolites on its surface for surveillance by MR1-specific immune cells, namely mucosal associated invariant T (MAIT) cells.
MR1 is known to be important for the recognition of microbial infections, the identification of cancerous cells as opposed to healthy ones, and the regulation of autoimmune disease. Recent studies also suggest that MR1 can display a much broader range of molecules, including cancer-specific metabolites, which could serve as internal sensors for disease. Additionally, MR1 does not vary across the human population, making it a promising potential immunotherapy target. However, prior to this study, the molecular mechanics of MR1 were unknown—specifically, the role of chaperone molecules in recognizing and stabilizing MR1 molecules.
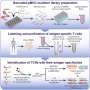
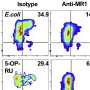
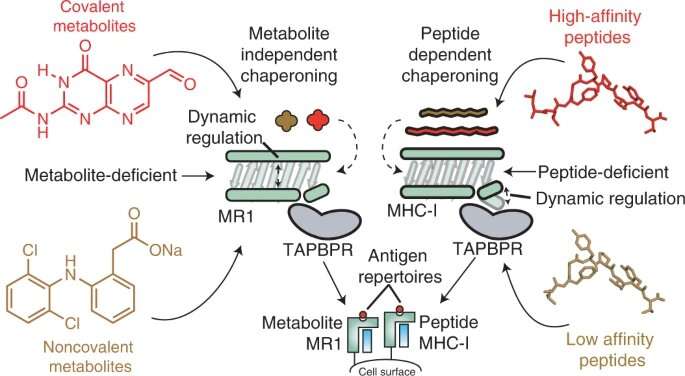
Using in vitro and in cell techniques, the researchers found that MR1 operates differently than MHC-I, particularly when it comes to molecular loading and affinity for the molecular chaperones Tapasin and TAPBPR. Whereas MHC-1 demonstrated a high affinity for molecular chaperones when empty but low affinity when loaded with presenting antigens, MR1 interacted at equally low levels with TAPBPR whether it was loaded with metabolites or not. Despite the short lifetime of the complex, researchers used NMR to successfully determine its 3D structure in a physiologically relevant, aqueous environment. Interestingly, these low-affinity interactions still enabled chaperones to catalyze loading and exchange of metabolite molecules directly on the MR1 surface.
"Our results provide a basis for utilizing molecular chaperones to stabilize MR1 molecules and to enable loading and presentation of antigens directly on the cell surface," said Nikolaos G. Sgourakis, Ph.D., Associate Professor in the Center for Computational and Genomic Medicine at Children's Hospital of Philadelphia and senior author of the study. "Future research should screen libraries of metabolites to see which show high affinity for binding to MR1. Researchers can then consider ways to increase MR1 surface expression, potentially by promoting more stable MR1/chaperone interactions via several approaches, such as the use of small molecules."
More information: Nikolaos Sgourakis, TAPBPR employs a ligand-independent docking mechanism to chaperone MR1 molecules, Nature Chemical Biology (2022). DOI: 10.1038/s41589-022-01049-9. www.nature.com/articles/s41589-022-01049-9
Journal information: Nature Chemical Biology
Provided by Children's Hospital of Philadelphia