Cryogenic electron microscopy reveals drug targets against common fungus
Most people carry the fungus Candida albicans on their bodies without it causing many problems. However, a systemic infection with this fungus is dangerous and difficult to treat. Few antimicrobials are effective, and drug resistance is increasing. An international group of scientists, including Albert Guskov, associate professor at the University of Groningen, have used single-particle cryogenic electron microscopy to determine the structure of the fungal ribosome. Their results, which were published in Science Advances on 25 May, reveal a potential target for new drugs.
Candida albicans usually causes no problems, or just an itchy skin infection that is easily treated. However, in rare cases, it may cause systemic infections that can be fatal. Existing antifungal drugs cause a lot of side effects and are expensive. Furthermore, C. albicans is becoming more drug-resistant, so there is a real need for new drug targets. "We noted that no antifungal drugs are targeting protein synthesis, while half of the antibacterial drugs interfere with this system," says Guskov. A reason for this is that fungal ribosomes, the cellular machineries that translate the genetic code into proteins, are very similar in humans and fungi. "So, you would need a very selective drug to avoid killing our own cells."
Atomic resolution
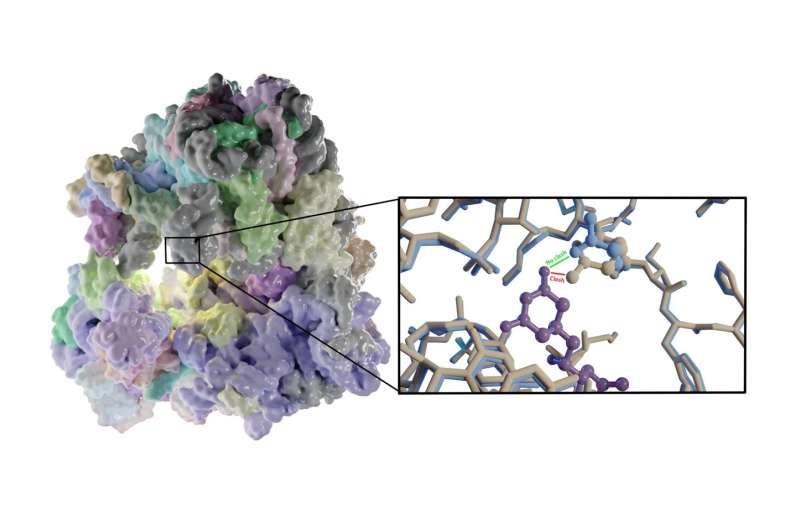
Therefore, Guskov and his collaborators reasoned that obtaining the structure of the C. albicans ribosomes would be valuable in finding drug targets. The classical approach is to grow crystals from purified ribosomes and to determine their structure using X-ray crystallography; however, this is a laborious technique. Instead, they used single-particle cryogenic electron microscopy, where a large number of single particles are imaged at very low temperatures in an electron microscope. The images of single particles—seen from different angles—are subsequently combined to produce an atomic-resolution structure.
Mutation
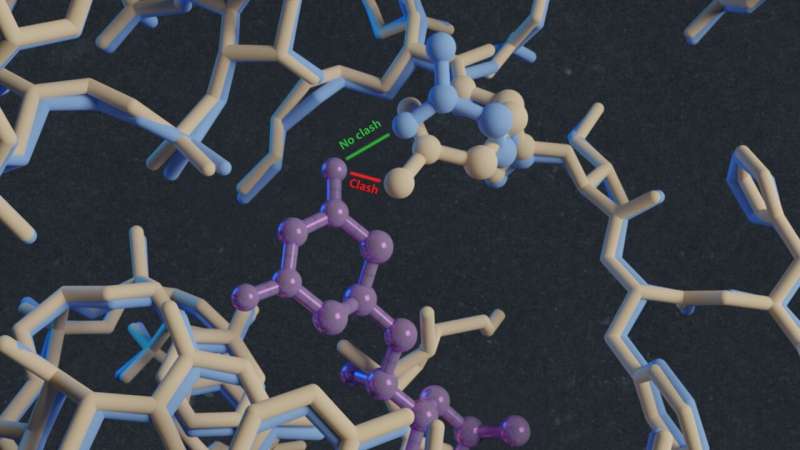
"In this way, we solved the structures of vacant and inhibitor-bound fungal ribosomes and compared their functions to those of ribosomes from yeast and rabbit—the latter as a model for the human ribosome—and repeated this for ribosomes bound to different inhibitors," explains Guskov. One of these inhibitors was the antimicrobial cycloheximide (CHX), to which C. albicans is known to be resistant. By comparing the structures, the scientists noted that a single mutation in the E-site, which plays a key part in protein synthesis, prevents CHX from binding to C. albicans ribosomes. "The mutation changed one amino acid in the structure of this E-site from proline to glutamine. This substitution reduces the size of the binding site, so the inhibitor can't attach and is therefore ineffective," notes Guskov. Another inhibitor, phyllanthoside, is not blocked by the mutation.
Threat
"By comparing the structures of the E-sites in vacant ribosomes in C. albicans and humans, and information on the way that different inhibitors bind to the site, we can develop a specific inhibitor that blocks fungal ribosomes but not those of humans. This would then be a selective drug to treat fungal infections," observes Guskov. The scientists are currently screening libraries of molecules to find drug leads. "It is extremely challenging to develop a vaccine against C. albicans, like we did for the coronavirus. So, we need drugs to treat systemic infections," he explains. "The increasing drug resistance of this fungus is a real threat. If this continues, we could be in serious trouble unless new drugs are developed."
More information: Yury Zgadzay et al, E-site drug specificity of the human pathogen Candida albicans ribosome, Science Advances (2022). DOI: 10.1126/sciadv.abn1062
Journal information: Science Advances
Provided by University of Groningen