Exploring ancient tuberculosis transmission chains
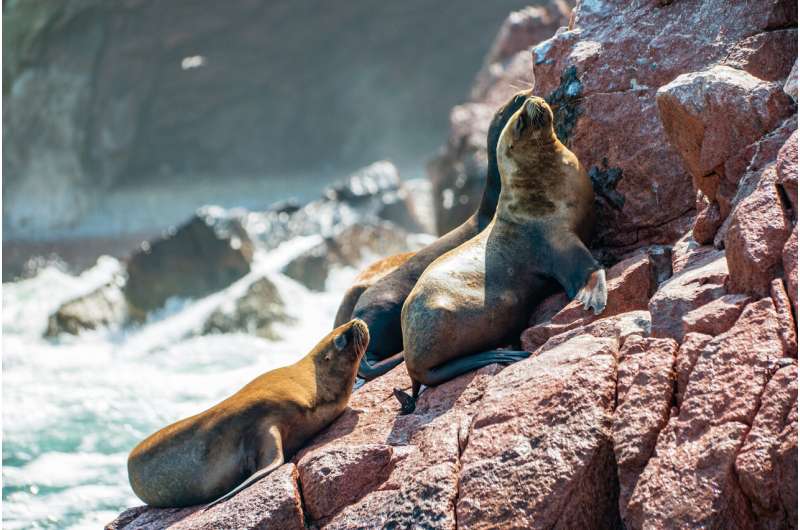

Tuberculosis (TB) is the second most common cause of death worldwide by an infectious pathogen (after Covid-19), but many aspects of its long history with humans remain controversial. Researchers at the Max Planck Institute for Evolutionary Anthropology in Leipzig, Germany, and Arizona State University in Tempe, USA, found that ancient TB discovered in archeological human remains from South America is most closely related to a variant of TB associated today with seals, but surprisingly these cases were found in people who lived nowhere near the coast. This implies that these cases were not the result of direct transmission from seals, and instead one, or more, spillover events were likely to be the primary drivers of human infection.
Nearly one quarter of the world's population is suspected to have been exposed to the bacterium responsible for tuberculosis, a disease that accounts for the highest global mortality from a bacterial infection. TB's global distribution was once viewed as support for its emergence deep in our past, where it was thought to have evolved in Africa tens of thousands of years ago and became distributed throughout the world following migrations with its host. Its ability to infect a number of mammalian species also make it a highly adaptable pathogen.
Analyses of ancient TB genomes have stirred up controversy about when this host-pathogen association began and precisely how TB became globally distributed. A 2014 study led by research teams at the University of Tübingen and Arizona State University reported on three ancient TB genomes from coastal Peru, which revealed aspects of its history that were incompatible with prevailing assumptions on TB's origins.
First, rather than identifying one of the well-characterized human-associated strains of the pathogen, the team identified a comparatively rare strain that today infects mostly marine mammals such as seals and sea lions (pinnipeds). In addition, their data suggested that TB was a much younger disease than previously thought, having emerged only sometime in the last 6000 years. "At the time, we assumed that TB made its way from Africa to the Peruvian coast through travel with infected seal populations," comments Kirsten Bos of the Max Planck Institute for Evolutionary Anthropology who co-led the new study. "We assumed the source of the infection in Peru had been a zoonosis from seals. It was not clear, though, if the specific TB infection we identified in the three people was a local phenomenon restricted to the area, or whether its distribution was broader".

TB is an infection well known to specialists in bone lesions and pathology. Paleopathologist Jane Buikstra of Arizona State University has extensively studied human skeletal remains across the Americas, and clear cases of TB infection are easily identified across the continents in the pre-contact period. "We've known for decades that a form of TB infection was present in the western coast of South America through the study of human remains. Now, with 21st century scientific advances, ancient DNA is the best tool available to investigate the relationships between the TB manifestations we observe osteologically in different parts of the Americas".
In a study published this week in Nature Communications, the team reports on three new cases of pre-contact era South American TB, this time from human remains that come from inland archeological sites, two of which are situated in the highlands of the Colombian Andes. All three people were infected with the marine-associated strain of TB, thus making a simple zoonosis from seals unlikely.
TB's entry into South America through human exposure to infected seals is still the strongest hypothesis, but how TB was subsequently distributed on land remains an open question. Lead authors Åshild Vågene and Tanvi Honap are confident that these new cases present strong evidence that the TB variant currently found in seals was once able to travel long distances on land. "The TB bacterium can infect numerous mammalian species, so there are many candidates for its terrestrial dispersal, including humans themselves," says Vågene. "Vast trade networks may have played an important role in transporting the pathogen from the coast". Honap adds that "recovering ancient TB DNA in animal remains from the pre-contact era Americas may one day allow us to explore the transmission chains responsible for bringing this marine variant so far inland".
Anne Stone of Arizona State University who specializes in the evolutionary history of TB and co-led the current study, sees the new results as an opportunity for deeper exploration into the ecology of the disease in the Americas before the colonial period. "It's an exciting time in ancient DNA research, as we can now look at genome-level differences in these ancient pathogens and track their movements across continents and beyond. For TB, the open question is how widespread the seal-associated strain was in human populations of the Americas prior to its replacement by the more virulent strains that arrived with the Europeans."
More information: Åshild J. Vågene et al, Geographically dispersed zoonotic tuberculosis in pre-contact South American human populations, Nature Communications (2022). DOI: 10.1038/s41467-022-28562-8
Provided by Max Planck Society