Insights into the past of an almost eradicated childhood infection
Haemophilus influenzae serotype b (Hib) was the main cause of bacterial meningitis in children and a major cause of worldwide infant mortality before the introduction of a vaccine in the 1980s.
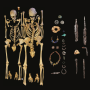
In the new study published in Genome Biology, led by the University of Tartu and the University of Cambridge, Guellil et al. reconstructed the oldest known and first ancient DNA Hib genome from a young Anglo-Saxon child, from 6th-century England, who probably died from a plague co-infection. A synthesis of genomic analyses and examination of the skeleton enabled the reconstruction of the genome of a bacterial pathogen often restricted to the respiratory tract, but able to cause severe infections of joints and the meninges as seen in this case.
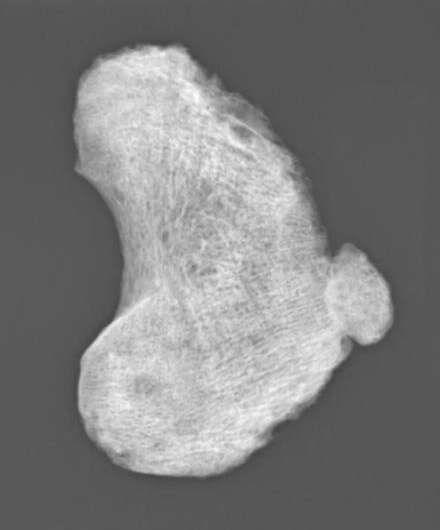
"The skeleton of the young boy (~6 years old) showed signs of disease consistent with bacterial septic arthritis, a progressive destruction of joints, probably caused by a prolonged and untreated Hib infection, which could have led to physical impairment and in the case of an additional meningeal infection also neurological impairment," said Guellil, the Research Fellow of Ancient DNA.
The additional recovery of a partial plague genome from the child also highlights how plague was affecting sub-adult populations during the historical pandemics. "Furthermore, it opens questions on the effect of plague infections in individuals with a weakened immune system already affected by infectious diseases," added Guellil.
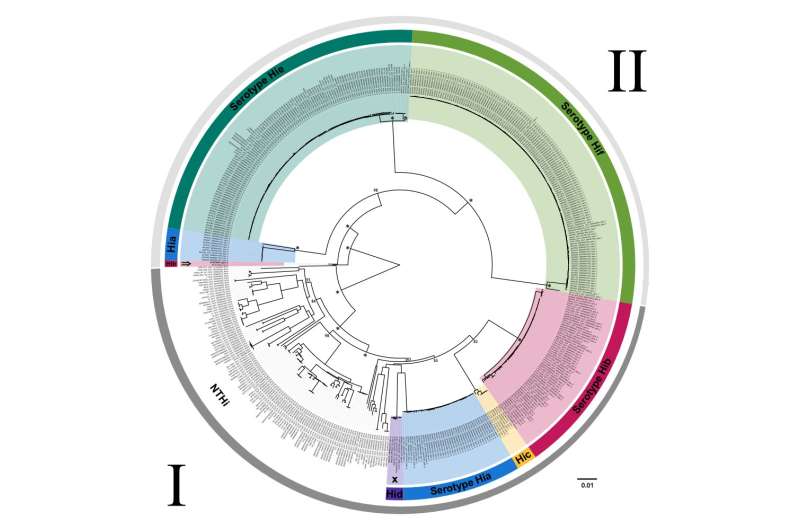
The analysis of the Hib genomes allowed for the first evolutionary insights into this major human pathogen and the origin of its b-capsule, which plays a major role in the virulence of the pathogen and is key to the current Hib vaccine. It also confirmed the presence of the pathogen with a similar clinical phenotype observed in the 20th century as early as the 6th century CE.The genome itself has a distinct virulence profile from current Hib genomes and places in the now ostensibly no longer circulating serotype b phylogroup. The similarity of our genome to the only other representative of this group, isolated in the 1940s, points to mostly clonal evolution of the clade since the 6th century CE.
Thanks to widespread vaccination programs, once common and potentially fatal Hib infections are now very rare, therefore the recovery of ancient genomes from skeletal remains offer unique opportunities to study the evolution of this pathogen
More information: Meriam Guellil et al, An invasive Haemophilus influenzae serotype b infection in an Anglo-Saxon plague victim, Genome Biology (2022). DOI: 10.1186/s13059-021-02580-z
Journal information: Genome Biology
Provided by Estonian Research Council