January 7, 2022 feature
Visualizing intracellular nanostructures of living cells using nanoendoscopy-AFM
Atomic force microscopy (AFM) offers a method for label-free imaging of nanoscale biomolecular dynamics to solve biological questions that cannot be addressed via other bioimaging methods including fluorescence and scanning electron microscopy. Since such imaging methods are only possible for biological systems extracted from cells or reconstructed on solid substrates, nanodynamics within living cells largely remain inaccessible with existing bioimaging methods. In a new report now published in Science Advances, Marcos Penedo and a research team in Nanolife Science and biotechnology at the Kanazawa University in Japan, overcame the limits of bioimaging by using nanoendoscopy-AFM. During the process, they inserted a needle-like probe into a living cell to present actin fiber, three-dimensional (3D) maps and 2D nanodynamics of the inner scaffold of the membrane with undetectable changes in cell viability. Unlike earlier AFM methods, the nanoprobe directly accessed the target intracellular components and explored the capabilities of AFM, including high-resolution imaging, nanomechanical mapping and molecular recognition to expand the observable range of intracellular structures in living cells.
Bioimaging intracellular dynamics
Molecular-scale dynamics of intracellular components provide insight to the fundamental mechanisms of cell functions and disease. However, direct imaging methods for such nanodynamics in living cells is challenging. For instance, while electron microscopy is useful to image nanostructures of frozen cells in vacuum, they are incapable of imaging nanodynamics in living cells under physiological environments, except as static snapshots of fixed conformations. Similarly, while fluorescence microscopy via fluorescence labeling provides a powerful method to visualize the dynamics of proteins and organelles in living cells, they are limited by an inability to efficiently image at the nanoscale. Strong demands therefore exist for a label-free intracellular imaging method in liquid environments. Atomic force microscopy (AFM) is a potential candidate for the role with the capacity to image at the sub-nanometer scale to visualize nanodynamics of lipids, proteins and DNAs without labels. However, such images are not representative of biological systems as a result of extraction from a cell or reconstruction on a solid substrate in vitro. In this work, therefore, Penedo et al. proposed an AFM-based imaging method known as nanoendoscopy-AFM to observe nanodynamics inside living cells without labeling or breaking them apart.
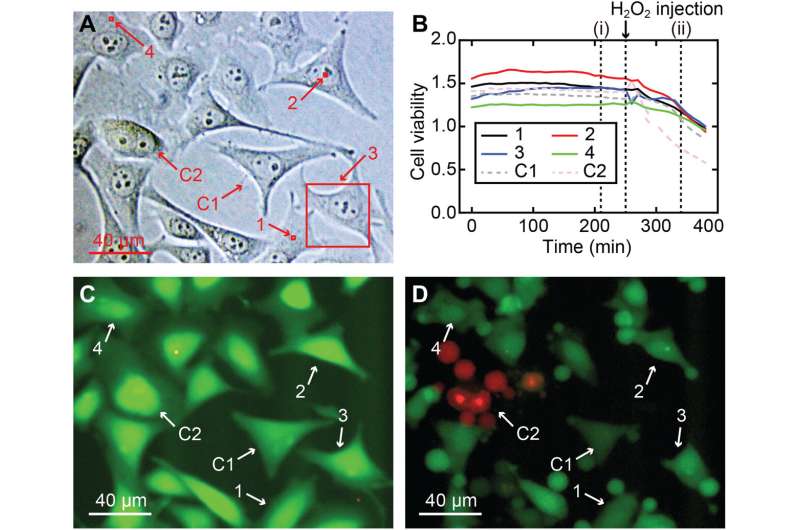
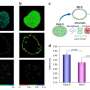
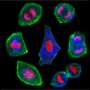
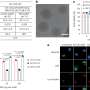
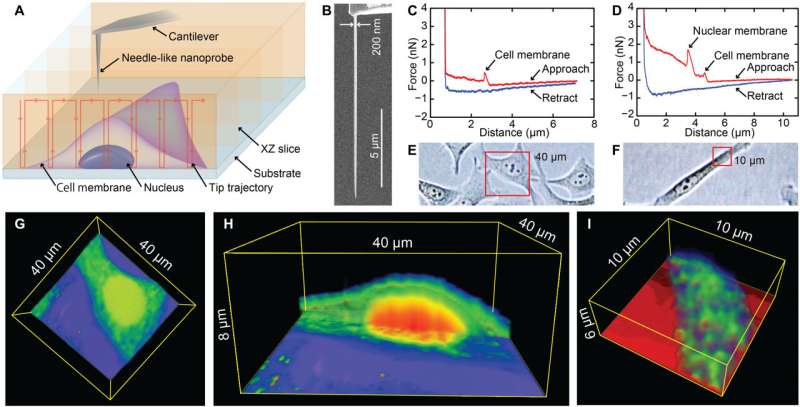
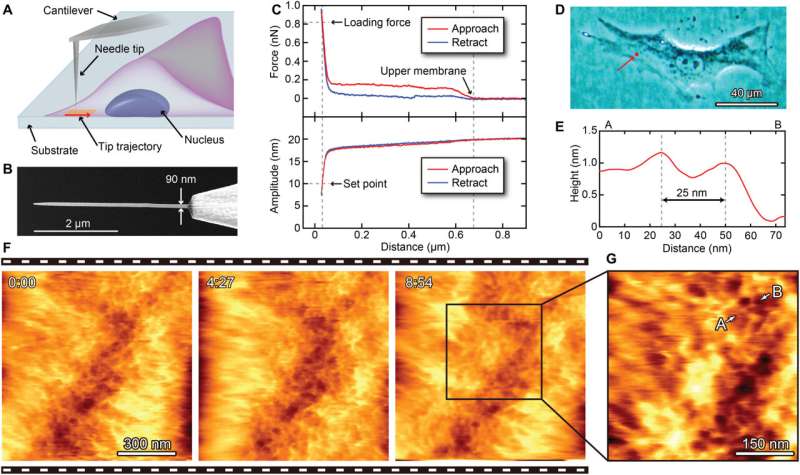
During the experiments, much like an endoscopic camera, the researchers inserted a long needle-like nanoprobe inside a living cell to perform 2D and 3D AFM imaging. The team showed how nanoendoscopy-AFM provided a unique advantage for label-free intracellular live-cell imaging at the nanoscale. The method provides a powerful path to observe hitherto unexplored phenomena in biological systems. Penedo et al. repeatedly introduced the nanoprobe inside the cell at different positions of the desired area via force versus distance curve measurements. In order to image the whole cell, the nanoprobe had to be long enough to completely penetrate the cell until it reached the substrate, and with diameters below 200 nm to minimize cell damage, while facilitating membrane penetration. The team used a commercial silicon tetrahedral tip as a nanoprobe, which they milled using focussed ion beam milling to the preferred dimensions. The team next used the nanoprobes inside different areas of a HeLa cell. They acquired a 3D nanoendoscopy-AFM image of a whole cell during the experiments and identified the nucleus of the HeLa cell from the rest of the cell. Further measurements also indicated the internal granular structures. To minimize cell damage during penetration, Penedo et al. reduced the penetration force and indentation length as much as possible. They also conducted cell viability experiments to confirm that 3D nanoendoscopy-AFM did not lead to severe cell damage when using nanoprobes with diameters below 200 nm. Using 3D nanoendoscopy-AFM, they facilitated imaging of the internal cytoskeleton in living cells to observe 3D organization of the unsupported fibers. The team also successfully merged intracellular images resulting from 3D nanoendoscopy-AFM and confocal microscopy.
2D nanoendoscopy-AFM
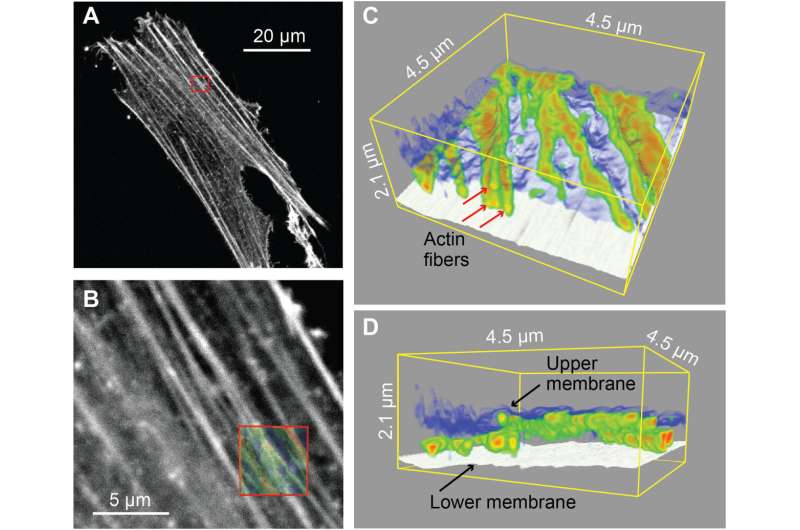
The ability to insert a long nanoprobe into a cell many times while maintaining cell viability implied the potential to locate the apex of the probe within a living cell to perform local 2D/3D AFM measurements without substantial damage. The nanoprobe could be inserted within the cell to measure the cytoplasmic side of the cell membrane via amplitude modulation mode AFM. The nanoprobes had to be long enough to completely penetrate the cell and reach its bottom, while being thin enough to reduce cell damage. To accomplish this in practice, Penedo et al. developed nanoprobes made of amorphous carbon using electron beam deposition and measured the amplitude dependence on the distance, to determine the integrity of the cell. They performed 2D nanoendoscopy-AFM experiments using a fibroblast cell to illustrate the reticular structure of the inner cell membrane and observed the cell architecture to study internal dynamics of cell structures. The work highlighted the possibility of using 2D nanoendoscopy-AFM to study nanodynamics of internal structures in living cells under physiological environments.
Outlook
In this way, Marcos Penedo and colleagues showed the applications of nanoendoscopy-AFM to measure cytoplasmic inner surfaces of cell membranes and associated scaffolds to understand 3D arrangement of actin filaments in their natural intracellular environment in living cells. The team sought to minimize cell damage by using ultrathin needle-like nanoprobes in the experiments. The proposed AFM methods produced 3D maps of internal cell structures in addition to 2D projections combined with existing fluorescence methods such as confocal or super-resolution microscopy. The method will shed light on cell machinery in action, in vivo, while exposing physiological molecular motors. The method will also open new possibilities to study intracellular nanomechanics that play an important role in cellular functions. The team can use the method to measure the stiffness, adhesion and dissipation characteristics of the nucleus to extract biological information suited for interdisciplinary fields of cell biology and medicine.
More information: Marcos Penedo et al, Visualizing intracellular nanostructures of living cells by nanoendoscopy-AFM, Science Advances (2021). DOI: 10.1126/sciadv.abj4990
Niels de Jonge et al, Electron microscopy of specimens in liquid, Nature Nanotechnology (2011). DOI: 10.1038/nnano.2011.161
Journal information: Science Advances , Nature Nanotechnology
Provided by Science X Network
© 2022 Science X Network