Researchers develop method for early detection of bacterial infection in crops
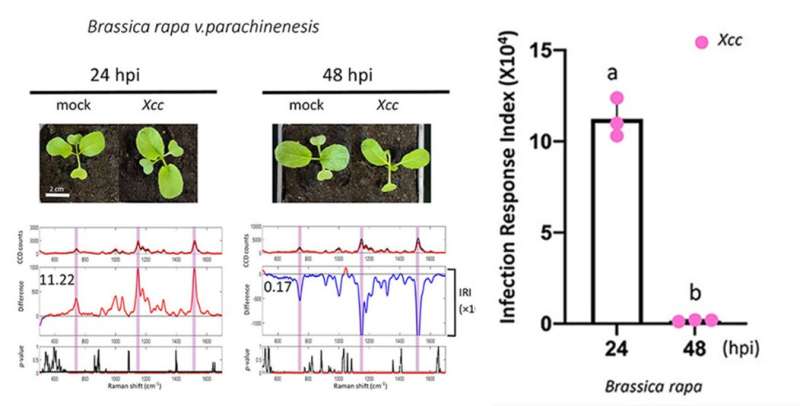
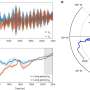
Researchers of Singapore-MIT Alliance for Research and Technology (SMART) and their local collaborators from Temasek Life Sciences Laboratory (TLL) have developed a rapid Raman spectroscopy-based method for the detection and quantification of early bacterial infection in crops. The Raman spectral biomarkers and diagnostic algorithm enable the non-invasive and early diagnosis of bacterial infections in crop plants, which can be critical for the progress of plant disease management and agricultural productivity.
Facing an increasing demand for global food supply and security, there is a growing need to improve agricultural production systems and increase crop productivity to overcome this challenge. Globally, bacterial pathogen infection in crop plants is one of the major contributors to agricultural yield losses. Climate change also adds to the problem by accelerating the spread of plant diseases. Hence, developing methods for rapid and early detection of pathogen-infected crops is important to improve plant disease management and reduce crop loss.
The breakthrough by SMART and TLL researchers offers a faster and more accurate method to detect bacterial infection in crop plants at an earlier stage, as compared to existing techniques. The team explained their research in a paper titled "Rapid detection and quantification of plant innate immunity response using Raman spectroscopy" published in Frontiers in Plant Science.
"The early detection of pathogen-infected crop plants is a significant step to improve plant disease management," says DiSTAP co-lead Principal Investigator Professor, TLL Deputy Chairman, and co-corresponding author, Chua Nam Hai. "It will allow the fast and selective removal of pathogen load and curb the further spread of disease to other neighboring crops."
Traditionally, plant disease diagnosis involves a simple visual inspection of plants for disease symptoms and severity. "Visual inspection methods are often ineffective as disease symptoms usually manifest only at relatively later stages of infection when the pathogen load is already high, and reparative measures are limited. Hence, new methods are required for rapid and early detection of bacterial infection. The idea would be akin to having medical tests to identify human diseases at an early stage, instead of waiting for visual symptoms to show so that early intervention or treatment can be applied," says DiSTAP Principal Investigator, MIT Professor, and co-corresponding author, Rajeev Ram.
While existing techniques, such as current molecular detection methods, can detect bacterial infection in plants, they are often limited in their use. Molecular detection methods largely depend on the availability of pathogen-specific gene sequences or antibodies to identify bacterial infection in crops; the implementation is also time-consuming and non-adaptable for on-site field application due to its high cost and bulky equipment required, making it impractical for use in agricultural farms.
"At DiSTAP, we have developed a quantitative Raman spectroscopy-based algorithm that can help farmers to identify bacterial infection rapidly. The developed diagnostic algorithm makes use of Raman spectral biomarkers and can be easily implemented in cloud-based computing and prediction platforms. It is more effective than existing techniques as it enables accurate identification and early detection of bacterial infection, both of which are crucial to saving crop plants that would otherwise be destroyed," explained Dr. Gajendra Pratap Singh, Scientific Director and Principal Investigator at DiSTAP, and co-lead author.
A portable Raman system can be used in agricultural farms and provides farmers with an accurate and simple yes or no response when used to test for the presence of bacterial infections in crop plants. The development of this rapid and non-invasive method will improve plant disease management and have a transformative impact on agricultural farms by efficiently reducing agricultural yield loss and increasing productivity.
"Using the diagnostic algorithm method, we experimented on several edible plants such as Choy Sum," says DiSTAP and TLL Principal Investigator and co-corresponding author Dr. Rajani Sarojam. "The results showed that the Raman spectroscopy-based method can swiftly detect and quantify innate immunity response in plants infected with bacterial pathogens. We believe that this technology will be beneficial for agricultural farms to increase their productivity by reducing their yield loss due to plant diseases."
The researchers are currently working on the development of high-throughput, custom-made portable or hand-held Raman spectrometers that will allow Raman spectral analysis to be quickly and easily performed on field-grown crops.
The development and discovery of the diagnostic algorithm and Raman spectral biomarkers were done by SMART and TLL. TLL also confirmed and validated the detection method through mutant plants.
More information: Pil Joong Chung et al, Rapid Detection and Quantification of Plant Innate Immunity Response Using Raman Spectroscopy, Frontiers in Plant Science (2021). DOI: 10.3389/fpls.2021.746586
Journal information: Frontiers in Plant Science