New group of antibacterial molecules identified
Researchers at Karolinska Institutet, Umeå University, and the University of Bonn have identified a new group of molecules that have an antibacterial effect against many antibiotic-resistant bacteria. Since the properties of the molecules can easily be altered chemically, the hope is to develop new, effective antibiotics with few side effects. The findings have been published in the scientific journal PNAS.
The increasing resistance to antibiotics in the world is alarming while few new types of antibiotics have been developed in the past 50 years. There is therefore a great need to find new antibacterial substances.
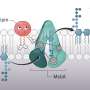
The majority of antibiotics in clinical use work by inhibiting the bacteria's ability to form a protective cell wall, causing the bacteria to crack (cell lysis). Besides the well-known penicillin, that inhibit enzymes building up the wall, newer antibiotics such as daptomycin or the recently discovered teixobactin bind to a special molecule, lipid II. Lipid II is needed by all bacteria to build up the cell wall. Antibiotics that bind to this cell wall building block are usually very large and complex molecules and therefore more difficult to improve with chemical methods. These molecules are in addition mostly inactive against a group of problematic bacteria, which are surrounded by an additional layer, the outer membrane, that hinders penetration of these antibacterials.
Attractive target for new antibiotics
"Lipid II is a very attractive target for new antibiotics. We have identified the first small antibacterial compounds that work by binding to this lipid molecule, and in our study, we found no resistant bacterial mutants, which is very promising," says Birgitta Henriques Normark, professor at the Department of Microbiology, Tumor and Cell Biology, Karolinska Institutet, and one of the article's three corresponding authors.
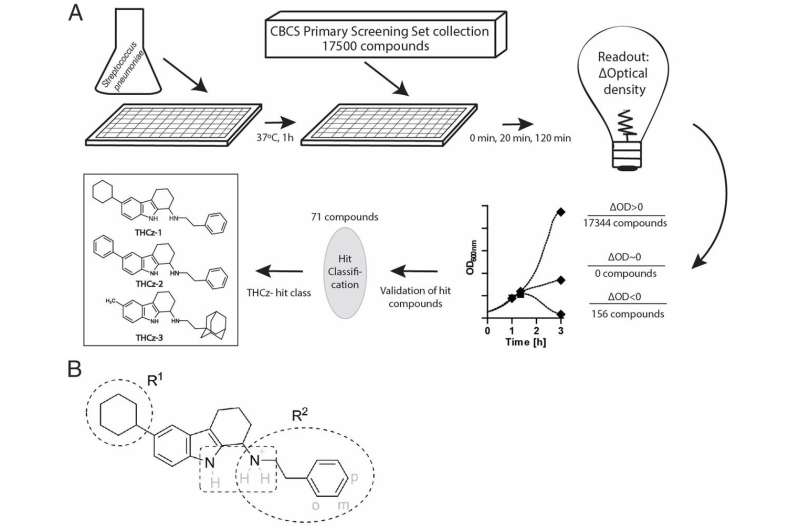
In this study, researchers at Karolinska Institutet and Umeå University in Sweden have tested a large number of chemical compounds for their ability to lyse pneumococci, bacteria that are the most common cause of community-acquired pneumonia. The initial tests were carried out in collaboration with the Chemical Biology Consortium Sweden (CBCS), a national research infrastructure at SciLifeLab. After a careful follow-up of active compounds from this screening, the researchers, in collaboration with the University of Bonn in Germany, found that a group of molecules called THCz inhibits the formation of the cell wall of the bacterium by binding to lipid II. The molecules could also prevent the formation of the sugar capsule that pneumococci need to escape the immune system and to cause disease.
More easy to change chemically
"The advantage of small molecules like these is that they are more easy to change chemically. We hope to be able to change THCz so that the antibacterial effect increases and any negative effects on human cells decrease," says Fredrik Almqvist, professor at the Department of Chemistry at Umeå University and one of the corresponding authors.
In laboratory experiments, THCz have an antibacterial effect against many antibiotic-resistant bacteria, such as methicillin-resistant staphylococci (MRSA), vancomycin-resistant enterococci (VRE), and penicillin-resistant pneumococci (PNSP). An antibacterial effect was also found against gonococci, which causes gonorrhea, and mycobacteria, bacteria that can cause severe diseases such as tuberculosis in humans. The researchers were unable to identify any bacteria that developed resistance to THCz in a laboratory environment.
"We will now also initiate attempts to change the THCz molecule, allowing it to penetrate the outer cell membrane found in some, especially intractable, multi-resistant bacteria," says Tanja Schneider, professor at the Institute of Pharmaceutical Microbiology at the University of Bonn and one of the corresponding authors.
More information: Elisabeth Reithuber et al, THCz: Small molecules with antimicrobial activity that block cell wall lipid intermediates, Proceedings of the National Academy of Sciences (2021). DOI: 10.1073/pnas.2108244118
Journal information: Proceedings of the National Academy of Sciences
Provided by Karolinska Institutet