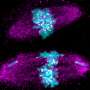
Experimentally reconstructing the reorganization of chromosomal DNA during mitosis
Two RIKEN researchers have discovered how, during cell division, one enzyme helps facilitate the tightly coordinated reorganization of chromosomal DNA into compact structures that can be evenly divided amongst the two resulting daughter cells1.
Ever since the dynamics of chromosomes undergoing mitosis—the process by which a cell splits into two identical daughter cells—were first described about 140 years ago, scientists have been trying to better understand the intricacies of this process.
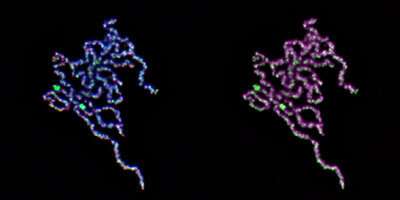
In the run-up to mitosis, each chromosome undergoes replication to form a pair of identical chromatids, with one chromatid slated for each daughter cell. By the time division begins, these two long chains of DNA have assembled into a pair of distinct, highly condensed structures, which are temporarily tethered to each other at their centers.
In 2015, Keishi Shintomi and Tatsuya Hirano, both of the RIKEN Chromosome Dynamics Laboratory, developed an assay that allowed them to induce chromatid assembly by combining isolated sperm nuclei with a mixture of six proteins. "Reconstituting a cellular process in a test tube using purified components is one of the most powerful approaches for dissecting and comprehending that process," explains Hirano.
While this technique has given valuable insights into the early stages of mitosis, critical elements of the chromatid assembly process remained poorly understood. In particular, Shintomi and Hirano sought to discover the role of topoisomerase IIα—one of the six proteins required for this assay—and set about optimizing their method in order to better characterize the function of this enzyme.
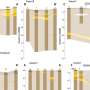
Their experiments revealed that topoisomerase IIα plays two roles in chromatid assembly. "The first is to dissolve entanglements between chromatids," says Shintomi. "While the second is to generate intra-chromatid entanglements that support condensation."
The disentangling function of topoisomerase IIα has been reported before, but the latter activity was unexpected. It appears to occur only when the enzyme is in a highly crowded environment with closely packed DNA strands. The two researchers were also able to identify a critical domain in topoisomerase IIα that directly coordinates the 'knotting up' of DNA during the chromatid condensation process.
Shintomi and Hirano are now keen to characterize other molecules that interact with topoisomerase IIα as part of the condensation process, including a class of DNA-binding proteins called linker histones. These proteins were absent in the researchers' original chromatid assembly assay, but the present study indicates that they interact with the same chromosomal regions that ultimately get targeted for proper actions of topoisomerase IIα.
"Such efforts will shed new light on the previously unrecognized action of linker histones and fierce competition among chromosomal proteins in the crowded environments of mitotic chromosome assembly," says Hirano.
The research was published in Nature Communications.
More information: Keishi Shintomi et al, Guiding functions of the C-terminal domain of topoisomerase IIα advance mitotic chromosome assembly, Nature Communications (2021). DOI: 10.1038/s41467-021-23205-w
Journal information: Nature Communications
Provided by RIKEN