Omitting delays from outbreak models grossly underestimates epidemic severity
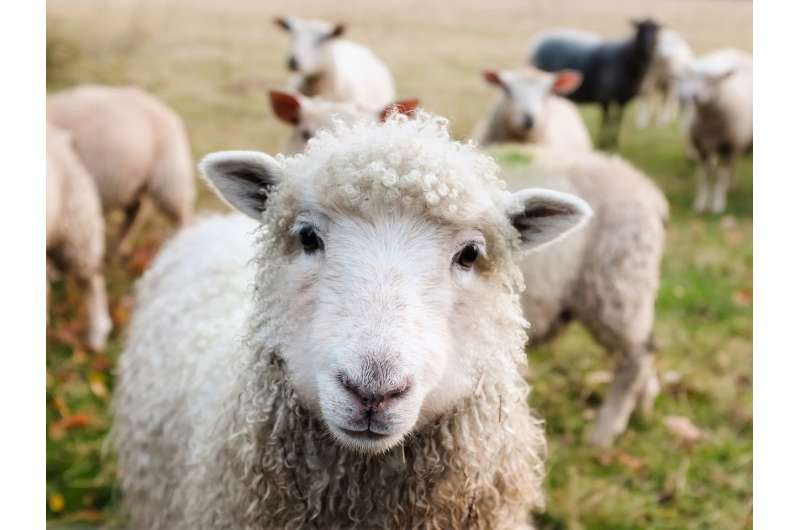

For livestock diseases, like foot-and-mouth disease (FMD) and swine flu, rapid culling and carcass disposal are well-established strategies for halting an outbreak and limiting its impact. However, even when infection is quickly detected delays in these interventions may permit pathogen transmission from infected farms.
A team of researchers has discovered that by neglecting to include response delays in their analyses, modelers grossly underestimate epidemic severity and its long-term consequences. The findings could also have implications for human diseases, including COVID-19.
"Livestock disease outbreaks can have devastating consequences economically, socially and politically," said Matthew Ferrari, associate professor of biology. "For example, preventing FMD outbreaks incurs an annual cost of $1.5 billion worldwide in FMD-free countries and an order of magnitude more in countries where the disease is endemic. This cost could be dramatically reduced if diseased herds are quickly detected and removed to prevent transmission to other animals."
Using the 2001 FMD outbreak in the United Kingdom as a case study, Ferrari, along with lead author Yun Tao, a former postdoctoral fellow at Penn State who is now an Intelligence Community Postdoctoral Research Fellow at the University of California Santa Barbara, and their colleagues, created a model to simulate how different response times would have affected the overall outbreak outcome.
"The United Kingdom government set ambitious targets for preventing the spread of FMD, yet there were still significant response lags that exacerbated the outbreak outcomes," said Ferrari.
The researchers used individual farm records collected by the United Kingdom Department for Environment, Food and Rural Affairs during the outbreak to explore the effects of three factors—farm size, control demand and farm density—on response delays. They defined farm size as the number of livestock on a farm to be culled, control demand as the number of farms scheduled for control and farm density as the number of infected farms within a geographical radius of 5 kilometers. Next, they simulated different delay times within a variety of contexts, including infected farms that were not yet culled and farms that were culled but with carcasses remaining.
"Our results demonstrated that farm size and control demand were key factors correlated with culling and disposal activities on individual farms," said Tao. "Specifically, veterinary response teams took longer to initiate responses on larger farms, which are major sources of potential spread. They also took longer when the outbreak was at its worst, likely because the system was overburdened."
For farm size, the team's model predicted that a farm that was larger by 100 animals was culled 3.7% slower, and the carcasses disposed of 2.2% slower. The number of farms that were waiting in the queue to be culled also correlated with drops in both culling and disposal efficiencies. For every 10 farms in the queue, the daily culling rate was reduced by 13% and the daily disposal rate by 8.4%.
The findings published March 3 in the Journal of The Royal Society Interface.
"Our results suggest that models that assume fixed, timely responses grossly underestimate epidemic severity and its long-term consequences," said Tao. "Including response dynamics and recognition of partial controllability of interventions in our models can help inform management priorities during epidemics of livestock diseases."
Ferrari added that the findings could have also implications for modeling human infectious diseases. "For a variety of reasons, we have seen healthcare and testing delays during the COVID-19 pandemic. Recognizing how operational delays may be exacerbated by the outbreak itself is a first step to developing robust strategies that could reduce the number of people who become sick in an outbreak."
More information: Yun Tao et al, Causes of delayed outbreak responses and their impacts on epidemic spread, Journal of The Royal Society Interface (2021). DOI: 10.1098/rsif.2020.0933
Journal information: Journal of the Royal Society Interface
Provided by Pennsylvania State University