Understanding oxygen-reducing enzymes

Methane is a powerful greenhouse gas that plays a central role in the global carbon cycle. At the same time, it is an important energy source for us humans. About half of its annual production is made by microorganisms known as methanogens that decompose organic material such as dead plants. This normally takes place in a habitat without oxygen as this gas is lethal to methanogens. But even in actually oxygen-free habitats, oxygen molecules occasionally appear. To render these intruders harmless, methanogens possess a special enzyme that is able to convert oxygen into water.
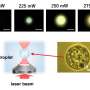
"Enzymes are vital components of the metabolism of all living organisms and the goal of our laboratory is to understand how these nanomachines are working at the molecular level," says Tristan Wagner from the Max Planck Institute for Marine Microbiology and first author of the study, published in the scientific journal Chemical Communication in September 2020. For the study, Wagner cultivated anaerobic microorganism called Methanothermococcus thermolithotrophicus, which originated from the sediment of the Gulf of Naples. He purified the enzyme F420-oxidase, a flavodiiron protein, and crystallized it, a common method to study the functioning of enzymes.
"It was already known that F420-oxidase can convert oxygen into water," says Wagner. "But we succeeded to decrypt the mechanism." The study is a cooperation of scientists from the Max Planck Institute for Marine Microbiology, the Max Planck Institute for Terrestrial Microbiology, the Paul Scherrer Institute, the Interdisciplinary Research Institute of Grenoble and the European Synchrotron Radiation Facility.
Oxygen is locked in
The mechanism, the researchers revealed, has an important requirement: Oxygen is very reactive, so it is crucial that the reaction is controlled correctly by the enzyme and no solvents are floating around. Otherwise the oxygen could accidentally be transformed in superoxide and kill the anaerobe. The trick of the enzyme F420-oxidase is to use a gas channel and a gating system. The oxygen molecule is first funneled in the specific channel to an appropriate anhydrous catalytic cavity containing iron. Then iron transforms the oxygen in water that will be released by a gating mechanism. For that the cavity begins to move and opens a small "door." Thanks to the movement, the newly generated water is transported outside. The empty cavity closes again and is available for the next oxygen molecule.
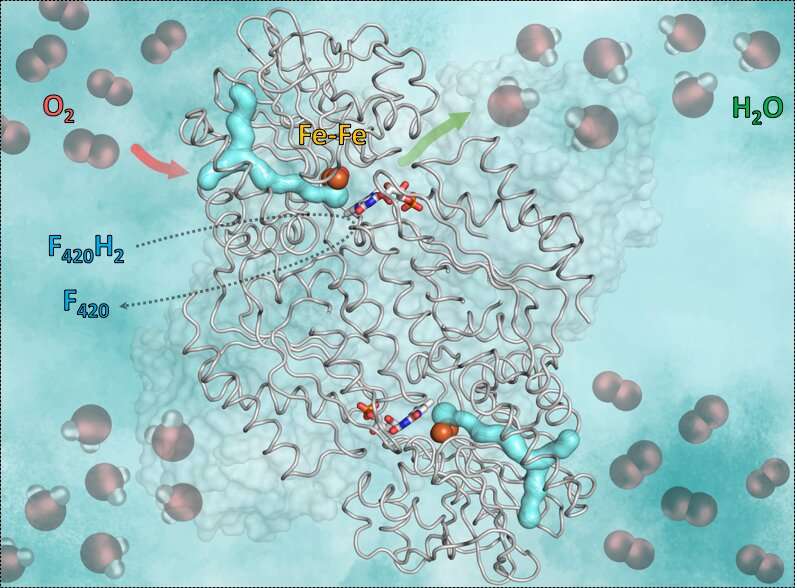
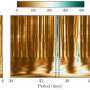
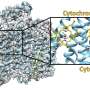
To gain insights into this mechanism the scientists used X-ray crystallography. They first obtained the crystal structure without oxygen, where they could see the anhydrous catalytic cavity isolated from the solvent. Then, they gassed the enzyme crystals with the inert gas krypton, which, unlike oxygen, can be made visible by X-rays. Afterwards they irradiated the enzyme crystals and were able to detect krypton atoms showing the gas channel leading to the catalytic cavity. The flavodiiron protein and its channel is conserved not only in methanogens, but also in other microorganisms like clostridia (who live mainly in soil or in the digestive tract), in the sulfur bacteria Desulfovibrio gigas or even in the intestinal parasite Giardia intestinalis.
The faster the better
"This reaction is really fast," says Sylvain Engilberge from the Paul Scherrer Institute and first author of the study next to Tristan Wagner. "This velocity is also the high importance of our investigation." Similar enzymes like laccase are much slower. "For future application of bio-inspired electrochemical processes, we need to learn more from the chemical reaction, structure and function of different groups of oxygen-reducing enzymes," says Engilberge. It would also pave the way of protein engineering to convert a high-rate O2-detoxifier into an electron sink for industrial processes.
"Our next step would be to understand the diversity of flavodiiron protein," says Tristan Wagner. Some homologues are not targeting oxygen but the poisonous nitric oxide, their enzymes can discriminate between both gases with high specificity. But what is the selective filter? The gas channel? The environment of the catalytic cavity? "More studies have to be carried out to understand how the protein discriminates oxygen and nitric oxide," adds Wagner. With such knowledge, it would be for instance possible to predict from genomic information if a flavodiiron protein would be an oxygen or a nitric oxide scavenger.
More information: Sylvain Engilberge et al, Krypton-derivatization highlights O2-channeling in a four-electron reducing oxidase, Chemical Communications (2020). DOI: 10.1039/d0cc04557h
Journal information: Chemical Communications
Provided by Max Planck Society