Fantastic muscle proteins and where to find them

Researchers at the Max Delbrück Center for Molecular Medicine in the Helmholtz Association (MDC) developed a mouse model that enables them to look inside a working muscle and identify the proteins that allow the sarcomere to contract, relax, communicate its energy needs, and adapt to exercise. Specifically, they were able to map proteins in defined subregions of the sarcomere, starting from the "Z-disc," the boundary between neighboring sarcomeres. This in and of itself was a significant step forward in the study of striated muscle.
In the process, they made an unexpected discovery: myosin, one of the three main proteins that make up striated muscle fibers, appears to enter the Z-disc. Models of how myosin, actin and the elastic scaffold protein titin work together have largely ignored the possibility that myosin filaments penetrate the Z-disc structure. Only recently have German scientists theorized that they do, but no experimental evidence has validated the model, until now.
"This is going to be unexpected even for myosin researchers," says Professor Michael Gotthardt, who heads MDC's Neuromuscular and Cardiovascular Cell Biology Lab and led the research. "It gets to the very basics of how muscles generate force."
Who's there?
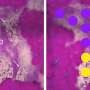
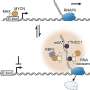
Gotthardt's team including first authors Dr. Franziska Rudolph and Dr. Claudia Fink with the help from colleagues at the MDC and the University of Göttingen, never set out to validate this theory. Their primary goal was to identify the proteins in and near the Z-disc. To do this, they developed a mouse model with an artificial enzyme, called BioID, inserted into the giant protein titin. The Titin-BioID then tagged proteins close to the Z-disc.
Sarcomeres are tiny molecular machines, packed with proteins that tightly interact. Until now it has been impossible to separate proteins specific to the different subregions, especially in live, functioning muscle. "Titin-BioID probes specific regions of the sarcomere structure in vivo," says Dr. Philipp Mertins, who heads MDC's Proteomics Lab. "This has not been possible before."
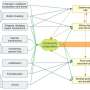
The team is the first to use BioID in live animals under physiological conditions and identified 450 proteins associated with the sarcomere, of which about half were already known. They found striking differences between heart and skeletal muscle, and adult versus neonatal mice, which relate to sarcomere structure, signaling and metabolism. These differences reflect the need of adult tissue to optimize performance and energy production versus growth and remodeling in neonatal tissue.
"We wanted to know who's there, know who the players are," Gotthardt says. "Most were expected, validating our approach."
The surprise
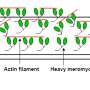
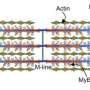
The protein that they were not expecting to see in the Z-disc was myosin, which is integrated at the opposite site of the sarcomere. When a muscle is triggered to move, myosin walks along actin bringing neighboring Z-discs closer together. This sliding of actin and myosin filaments creates the force that enables our heart to pump blood or our skeletal muscle to maintain posture, or lift an object.
This so-called "sliding filament model" of the sarcomere describes force production and helps explain how force and sarcomere length relate. However, current models have trouble predicting the behavior of fully contracted sarcomeres. Those models have assumed myosin does not enter the Z-disc on its walk along actin. There have been some hints that maybe it keeps going. "But we didn't know if what we were seeing in stained tissue samples was an artefact or real life," Gotthardt says. "With BioID we can sit at the Z-disc and watch myosin pass by."
Gotthardt agrees with the proposed theory that myosin entering the Z-disc can limit or dampen the contraction. This might help solve the ongoing issue scientists have had calculating how much force a muscle fiber can create in relation to its length and lead to a refined model of the sarcomere and possibly serve to protect muscle from excessive contraction.
Why it matters
Understanding how muscle fibers extend and contract on the molecular level under normal conditions is important so researchers can then identify what is going wrong when muscles are damaged, diseased or atrophy with age. Identifying which proteins are causing problems could potentially help identify novel treatment targets for patients with heart disease or skeletal muscle disorders.
Gotthardt and his team plan to next use BioID to study animals with different pathologies, to see what proteins are involved in muscle atrophy, for example. "Maybe a protein that is not normally there goes into the sarcomere, and it is part of the pathology," Gotthardt says. "We can find it with BioID."
More information: Franziska Rudolph et al, Deconstructing sarcomeric structure–function relations in titin-BioID knock-in mice, Nature Communications (2020). DOI: 10.1038/s41467-020-16929-8
Journal information: Nature Communications
Provided by Max Delbrück Center for Molecular Medicine