A new tool to map the flow of info within living cells
How do cells move? Why do they move? Why do some cancer cells move slowly while others move quickly, causing a cancerous tumor to metastasize and become much more difficult to treat effectively? The answers are not as simple as we would want. They involve tiny proteins and processes that are very difficult to study in real time and space. The UNC Department of Pharmacology labs of Klaus Hahn, Ph.D., and John Sondek, Ph.D., are dedicated to overcoming this difficulty. And now their labs, with researchers at the University of Texas Southwestern Medical Center, have created a way to study and map the intricacies of intercellular signaling—when, where, and how tiny parts of cells communicate—to make cells move.
Published in Nature Chemical Biology, the research provides a much-needed method for studying the precise movement mechanisms in healthy cells in real time and how these mechanisms might change in disease states, such as cancer metastasis.
"Our new tools allowed us to map the flow of signaling information within living cells and measure how much specific proteins contribute to cell behaviors, such as cell migration," said co-first author Daniel Marston, Ph.D., assistant professor in the UNC Department of Pharmacology at the UNC School of Medicine.
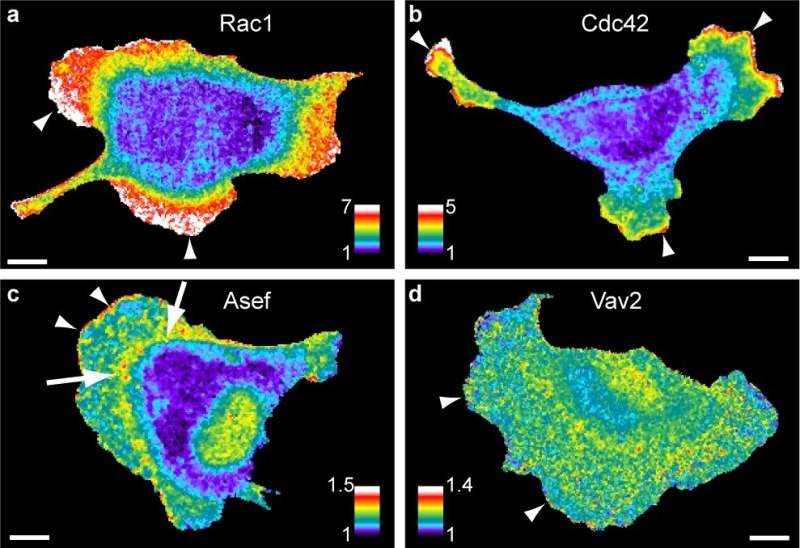
To do this, Marston and colleagues relied on microscopy tools developed in the lab of Klaus Hahn, the Ronald G. Thurman Distinguished Professor of Pharmacology. These microscopy tools, which use fluorescent biosensors, allowed the researchers to visualize the activity of multiple proteins in living cells at the same time. Then, thanks to mathematical analytic methods developed at UT Southwestern Medical Center, the research team quantified how proteins regulate each other.
Together, these tools can provide precise information about how signaling networks are wired together and how they regulate cellular processes, such as cell migration and metastasis. They will allow researchers at UNC and elsewhere to understand how these networks operate in healthy cells. With that information in hand, researchers could compare data from healthy cells to cell movement during various health conditions, such as inflammation—a hallmark of many diseases.
"If we can find out how these movement processes are changed in diseases, such as cancers, then we might be able to design better treatments effective only for that particular disease, while keeping other cells healthy," Marston added.
More information: Marston, D.J., Vilela, M., Huh, J. et al. Multiplexed GTPase and GEF biosensor imaging enables network connectivity analysis. Nat Chem Biol (2020). doi.org/10.1038/s41589-020-0542-9
Journal information: Nature Chemical Biology
Provided by University of North Carolina Health Care