How cellular machinery labels proteins for degradation
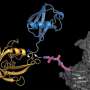
Proteins are molecular work horses in the cell that perform specific tasks, but it is essential that the timing of protein activities is exquisitely controlled. When proteins have fulfilled their tasks, their degradation ends processes that are unneeded or detrimental. To control timing, a label called ubiquitin is attached to unwanted proteins, marking them for degradation. Although complex molecular machineries were known to attach ubiquitin, how these machines carry out the labeling process was unknown. Researchers at MPIB, in collaboration with the University of Nevada Las Vegas have revealed these mechanisms and published the results in the journal Nature.
Numerous cellular processes such as immune responses or cell multiplication depend on many different proteins working in sequence. In order for the cell to function correctly, proteins must be degraded after their work is done. When disease-causing mutations block timely protein degradation, proteins could function at the wrong time, which can lead to diseases, including cancers, heart diseases and developmental disorders.
Controlling protein degradation
Cells break down proteins by marking unwanted proteins for degradation with another protein called ubiquitin. The labeling process, known as ubiquitylation, is carried out by molecular machines called E3 ligases. It is important that the E3 ligases themselves are switched on and off in the cells at the right place and at the right time. The on-switch for about one-third of all E3 ligases is a small protein that is similar to ubiquitin called NEDD8.
NEDD8 at the control of turning off other proteins
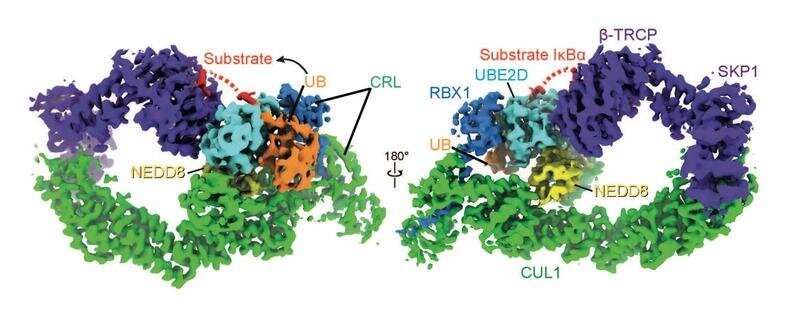
Although the individual components of these protein degradation machineries were known, it was unclear how NEDD8 switches on the E3 ligases and enables tagging the target protein with ubiquitin. "This is especially important because there are drugs in anti-cancer clinical trials that block NEDD8, and some infectious bacteria manipulate NEDD8 to disturb cellular processes," said Brenda Schulman, head of the Molecular Machines and Signaling department at MPIB. Schulman and her team have now deciphered the molecular mechanisms of this ubiquitylation. "We investigated the mode of action of an E3 turned on by NEDD8. We discovered how NEDD8 induces an E3 molecular machine to bring the ubiquitin tag to its targets. This is a key to switching proteins off at the right time when no longer needed in a cell," said Schulman.
Using chemistry and cryo-electron microscopy, the scientists have succeeded in visualizing an important E3 ligase activated by NEDD8 in the process of ubiquitylation. "To do this, we took a close look at each step in the tagging process. The natural process occurs within a fraction of a second, after which the molecular tagging machine falls apart. The process of capturing this normally short-lived state was particularly difficult," explains Kheewoong Baek, lead author of the study.
E3 ligase molecular machines control many cellular processes. "The deciphered mechanism not only explains the normal process and what goes wrong in some cancers where mutations prevent the E3 machine from working, but can also serve as a guide for developing therapies to tag unwanted proteins with ubiquitin. We hope that in the long-term this could help degrade proteins that cause cancer," says Schulman.
More information: Kheewoong Baek et al. NEDD8 nucleates a multivalent cullin–RING–UBE2D ubiquitin ligation assembly, Nature (2020). DOI: 10.1038/s41586-020-2000-y
Journal information: Nature
Provided by Max Planck Society