Expanding virophage diversity
Virophages are small viruses with double-stranded DNA genomes that co-infect eukaryotic cells along with giant viruses. Almost all known virophage genomes share only four genes in common: major and minor capsid proteins (MCP and mCP, respectively), ATPase involved in DNA packaging, and PRO, a cysteine protease involved in capsid maturation.
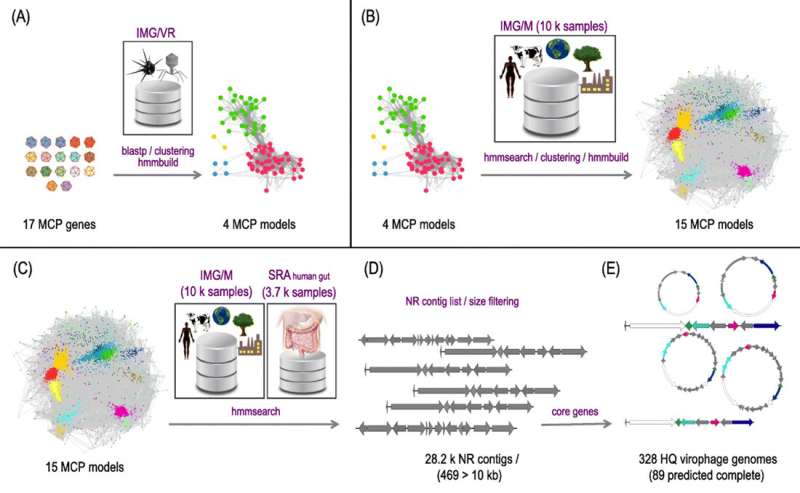
Recently reported in Microbiome, researchers from the US Department of Energy (DOE) Joint Genome Institute (JGI), a DOE Office of Science facility, have increased the number of known high quality virophage genome sequences 10-fold through computational approaches. By mining more than 14,000 publicly available metagenomic datasets in JGI's Integrated Microbial Genomes & Microbiomes (IMG/M) data suite—which includes IMG/VR (for Virus)—for the virophage marker gene MCP, they were able to identify 44,221 total virophage partial sequences, including over 28,000 unique MCP sequences.
Further analysis led to the identification of 328 "high quality" (based on completeness) diverse new virophage genomes containing all four core genes. These virophages were found in diverse habitats including the air, plant rhizosphere, wastewater, and even animal and human (for the first time) gut and taxonomically classified into 27 distinct clades (17 of them without previously known representatives). Of these, 89 contigs were considered to be complete genomes, and their discovery has extended the possible virophage genome size range from 13.8-29.3 kilobases (Kb) to 10.9-42.3 Kb. Additionally, the gene counts have similarly gone up from 13-25 to 12-39. "Overall," the team concluded, "we provide a global analysis of the diversity, distribution, and evolution of virophages."
More information: David Paez-Espino et al. Diversity, evolution, and classification of virophages uncovered through global metagenomics, Microbiome (2019). DOI: 10.1186/s40168-019-0768-5
Provided by DOE/Joint Genome Institute